-
沙棘(Hippophae rhamnoides L.)是中国重要的荒山荒坡造林、治理水土流失、改善生态环境的先锋树种[1],但干旱环境仍然限制着沙棘的生长和功能的发挥。叶片作为植株光合和蒸腾作用的重要器官,其表型变化是干旱胁迫下最为直观的重要特征。表观遗传修饰通过改变染色质状态和招募各种转录调节因子来控制基因的表达,从而影响植物对干旱胁迫等环境的响应[2]。组蛋白修饰包括组蛋白核心蛋白H2A、H2B、H3和H4的N末端特定氨基酸残基的甲基化、乙酰化、磷酸化和泛素化等[3]。组蛋白乙酰化参与染色质解缩和基因激活从而影响基因转录[4],作为一个动态可逆过程,组蛋白乙酰化和去乙酰化分别由组蛋白乙酰转移酶(HATs)和组蛋白脱乙酰酶(HDACs)催化。而HDACs已被证明在高粱(Sorghum bicolor L.)[5]、棉花(Gossypium hirsutum L.)[6]、拟南芥(Arabidopsis thaliana L.)[7]和大豆(Glycine max L.)[8]等植物中参与逆境响应和生物胁迫防御。有研究发现番茄(Solanum lycopersicum)组蛋白脱乙酰酶基因SlHDA3参与了番茄对干旱胁迫的响应,干旱胁迫和脱落酸处理显著诱导了SlHDA3的表达,沉默SlHDA3后植株对干旱胁迫的耐受性降低,干旱胁迫下脱落酸(ABA)、叶绿素、丙二醛(MDA)和脯氨酸(Pro)含量增加[9]。
曲古抑菌素A(trichostatin A, TSA) 是一种强效可逆的组蛋白去乙酰化酶抑制剂,可通过抑制HDAC而影响基因的表达[10]。研究发现TSA可以在干旱胁迫条件下显著上调花生(Arachis hypogaea L.)AhDREB1基因的表达,从而提高抗旱性[11]。在花生中组蛋白去乙酰化酶基因AhHDA1可以通过与其启动子结合来抑制AhGLK1的表达,TSA处理后AhGLK1的表达增加,H3乙酰化减少,证实AhGLK1受组蛋白去乙酰化修饰的调节[12]。在低温胁迫下,TSA处理强烈抑制了ZmDREB1和ZmCOR413等玉米(Zea mays Linn. Sp.)冷响应基因的表达[13]。另外,TSA还可以提高植物的抗寒性,并影响At1g55960、At3g50970和At5g57560等拟南芥冷诱导基因的表达[14]。
前期研究共筛选到沙棘ABF2、NAC2、C4H2、CHS4、HDA6和HDA19等干旱胁迫响应核心基因[15],揭示了以 ABA 依赖信号途径为主的信号传递途径和以类黄酮途径为主的氧自由基清除途径的耐旱分子机制[16],然而在外施TSA处理沙棘扦插苗响应干旱时,组蛋白去乙酰化酶基因表达水平的变化以及抗旱相关的ABA和类黄酮合成相关基因表达的影响不清楚。本研究探讨TSA预处理后沙棘叶片对干旱胁迫响应的表型和生理变化以及组蛋白乙酰化相关基因表达水平的变化,将为后续深入研究沙棘抗旱基因调控机制奠定基础。
-
本试验使用的材料采自中国林业科学研究院沙漠林业实验中心,将引进蒙古大果沙棘‘向阳’(H.rhamnoides L. subsp. mongolica, ‘Xiangyang’, XY) 2年生扦插苗种植于中国林业科学研究院科研温室大棚,生长条件为自然光照,昼/夜温度为20~30 ℃/10~15 ℃,相对湿度在60%~70%,正常生长2个月。在试验前一周将苗木移至恒温培养箱中,设定条件为:温度为(25 ± 1) ℃,光照度为(4 500 ± 500)LX,光照时间12 h,湿度70% ± 10%,使扦插苗稳定适应环境。
-
选取健康、生长均一的沙棘扦插苗带根部取出, 用1/2 Hoagland 营养液洗净土壤,洗净后的沙棘苗放在营养液中缓苗一周,之后在1 μmol·L−1 TSA(TSA, Solarbio)溶液中浸泡12 h,取部分叶片材料保存(单独TSA处理,T0),剩余沙棘苗浸泡于20%聚乙二醇(PEG6000, Solarbio)溶液中模拟土壤干旱胁迫, 分别在PEG处理12 h和48 h后取叶片材料保存(TSA + PEG复合处理,T12和T48),剩余植株取出后再分别浸泡于1/2 Hoagland 营养液正常培养3 d取叶片保存(复合处理后复水3 d处理,T12-3和T48-3)。以1/2 Hoagland 营养液正常培养植株为对照(CK)。每个分组处理均设置3次生物学重复。
-
将沙棘扦插苗带根取出洗净后在1/2 Hoagland 营养液中浸泡12 h, 再转移至20% PEG 6000溶液中, 分别在PEG处理12 h和48 h后取叶片材料保存(干旱胁迫处理,Y12和Y48),剩余植株取出后再分别浸泡于1/2 Hoagland 营养液正常培养3 d取叶片保存(干旱胁迫后复水3 d处理,Y12-3和Y48-3)。以1/2 Hoagland 营养液处理为对照(CK)。每个分组处理均设置3次生物学重复。
-
在每个干旱处理节点分别选取长势一致的幼苗 10 株,蒸馏水洗净滤纸吸干后用分析天平称量扦插苗鲜质量(TB)。在每个处理节点分别选取长势一致的幼苗 3株,用 LI-6400 便携式光合仪(LI-COR,美国)分别测定对照组与试验处理组沙棘幼苗叶片的净光合速率(Pn,μmol·m−2·s−1)、蒸腾速率(Tr,mmol·m−2s−1)和气孔导度(Cond,mol·m−2·s−1)。测定条件:红蓝固定光源,固定光强为1 000 μmol·m−2·s−1,温度 28 ℃,相对湿度为70%,二氧化碳浓度为温室中大气浓度。同时,使用Yaxin-1161G叶绿素荧光仪分别测量各处理幼苗叶片的PSⅡ最大光化学效率或原初光能转换效率Fv/Fm值和PSⅡ有效光化学量子产量[Y(Ⅱ)]值,使用SPAD502叶绿素含量测定仪测定叶片叶绿素相对含量(SPAD)。另外,使用脯氨酸(Pro)含量检测试剂盒(Solarbio)测定叶片游离脯氨酸含量,使用丙二醛(MDA)含量检测试剂盒(Solarbio)测定叶片丙二醛含量,使用植物脱落酸(ABA)酶联免疫分析试剂盒(Jingmeibio) 测定叶片ABA含量,使用植物类黄酮含量检测试剂盒(Solarbio)测定叶片类黄酮含量。每个分组处理均设置3次生物学重复。
-
使用RNAprep Pure多糖多酚植物总RNA提取试剂盒提取沙棘幼苗叶片RNA。使用PrimeScript RT试剂盒与gDNA Eraser试剂盒进行反转录。选择干旱胁迫过程中对以脱落酸依赖信号途径为主的信号传递途径有影响的ABA合成关键基因ABF2和NAC2,对以类黄酮途径为主的氧自由基清除途径有影响的类黄酮合成关键基因C4H2和CHS4,以及组蛋白去乙酰化酶基因HrHDA6和HrHDA19-1,利用Primer 5.0软件设计引物(表1)。使用StepOnePlus RT-PCR仪器进行实时荧光定量PCR反应。以18SrRNA基因为内参,扩增程序为95 ℃预变性30 s;95 ℃变性5 s, 60 ℃退火延伸34 s, 共40个循环。基因表达量使用2−△△CT方法进行分析[17]。
表 1 沙棘内参基因和目的基因引物
Table 1. Primers for reference genes and target genes in sea buckthorn
引物名称
Primer name正向引物
Forward primer(5′→3′)反向引物
Reverse primer(5′→3′)功能注释
Functional annotation18S rRNA GCCAAGGAAGTTTGAGGCAA TTCAAAGATTACCCGGGCCT 内参基因 Reference gene ABF2 TCCAGAACACGGTGGGTAGC AAGAGTCAACGAGCCTTGCCT 脱落酸合成途径
Abscisic acid synthesisNAC2 GCTCAAAAGCCGACGACAAG CGAATTCACGCGAGGCAAAT C4H2 AACCCGAAGAGTTCCGACCA CAAGAATAGGCAAAGCCAGAATGAT 类黄酮合成途径
Flavonoid synthesis pathwayCHS4 GTCAACCTAAGTCCAAAATAACCCA CGAAGAACTGTACCTCCAGCAAA HrHDA6 GGGGGGAGCATCTTTACCAT GCTTCATTGGGTGACCTTGG 组蛋白去乙酰化酶基因
Histone deacetylase geneHrHDA19-1 GACCTCCAGACACTGACATTCCA GCTCTCCTGATCTTTAACCTCGG -
使用IBM SPSS Statistics 24 统计软件中的单因素ANOVA分析方法,计算qRT-PCR数据获得P值,P<0.05表示差异具有显著性。使用Excel软件作图。
-
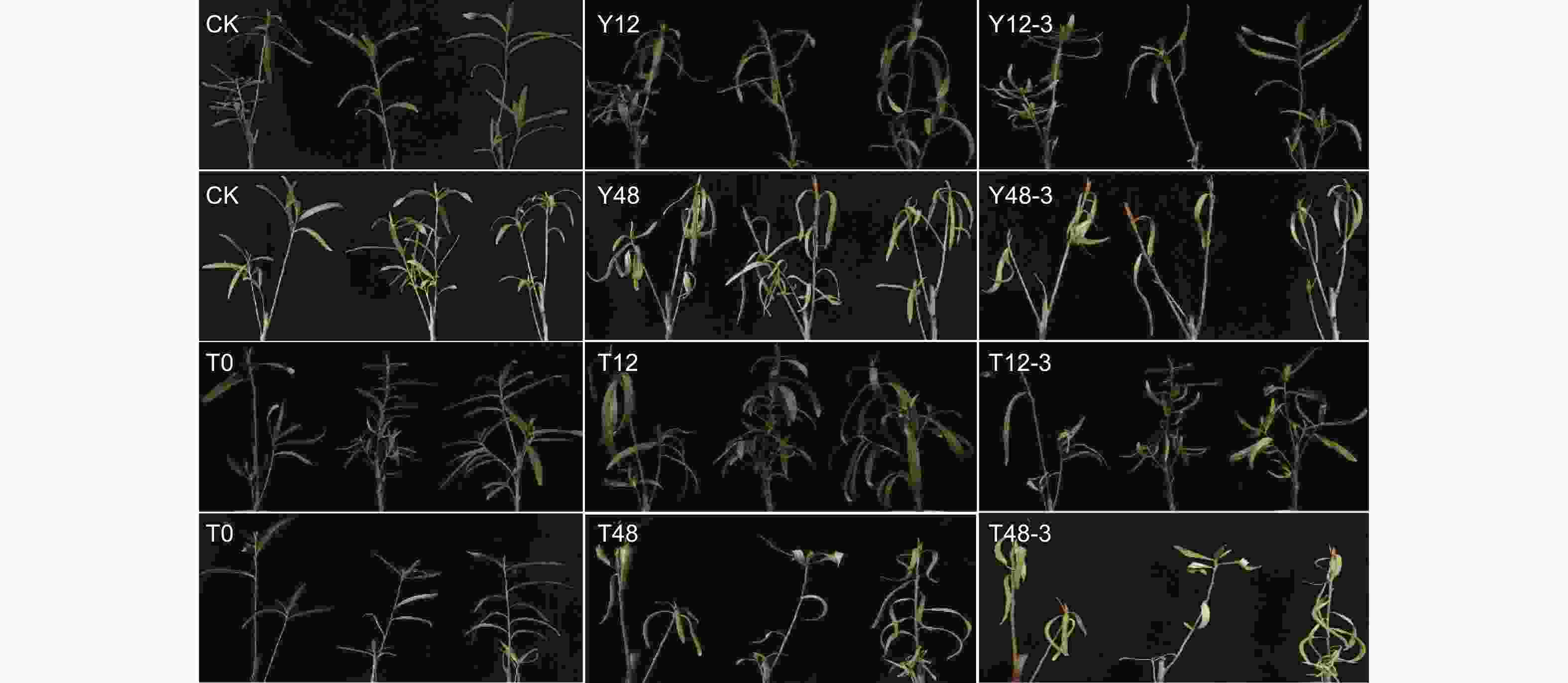
图1显示,与对照(CK)相比,单独干旱处理12 h后的沙棘植株叶片有下垂和萎蔫现象(Y12),在复水3 d后部分叶片恢复舒展(Y12-3),而干旱处理48 h后的沙棘植株叶片明显变黄且发生严重下垂和萎蔫现象(Y48),在复水3 d后仍无法恢复正常生长,后续发生叶片脱落(Y48-3);同时,经TSA预处理12 h再干旱处理后的沙棘植株的叶片下垂和萎蔫程度明显低于单独干旱处理的植株。与对照相比,干旱12 h后的植株鲜质量显著下调19.4%,经TSA预处理后的干旱12 h植株质量显著下调9.4%;干旱48 h后的植株质量显著下调38.6%,经TSA预处理后的干旱48 h植株质量显著下调21.3%(图2)。
-
干旱胁迫会影响沙棘光合生理特性、叶绿素荧光参数和叶绿素相对含量。在TSA预处理后的干旱处理过程中,净光合速率(Pn)随着干旱时间的增加而降低,干旱12 h后复水3 d,光合速率恢复明显,但干旱48 h后复水3 d时光合速率无明显变化(图3-A)。干旱显著影响沙棘的蒸腾速率(Tr),在干旱12 h后蒸腾速率显著下调83.0%,气孔导度(Cond)显著下调95.4%,干旱48 h后蒸腾速率显著下调98.7%,气孔导度显著下调99.0%,而在TSA预处理后干旱12 h时,蒸腾速率和气孔导度比干旱处理时显著提高(P<0.05),且复水后蒸腾速率和气孔导度部分恢复显著(图3-B, C)。

图 3 TSA预处理对沙棘干旱处理下和复水条件下叶片光合生理指标的变化
Figure 3. Effects of TSA on Photosynthetic physiological indexes of sea buckthorn under drought and rehydration
叶片叶绿素光系统II(PSII)接受光量子的最大能力Fm 与Fo的差值为光合作用的最大能力Fv,Fv/Fm是PSII的最大量子效率,Y(II)为实际光化学量子产量,二者可以作为叶绿素荧光参数衡量叶片光系统活性受损程度[18]。在干旱程度逐渐加深的过程中,Fv/Fm逐渐显著降低(图3-D)。在干旱12 h后Y(II) 显著下调22.0%,48 h后显著下调52.8%,而在TSA预处理后,Y(II)比干旱处理12 h时显著提高46.2%,比仅干旱处理48 h时显著提高62.8%,且复水后部分恢复显著(图3-E)。在干旱12 h后SPAD值显著下调12.3%,48 h后显著下调29.6%,而在TSA预处理后,SPAD值比仅干旱处理12 h时显著提高6.0%,比仅干旱处理48 h时显著提高11.0%(图3-F)。
-
沙棘受到干旱胁迫时体内脯氨酸含量明显升高,从而提高沙棘的渗透调节能力。与对照组相比,随着干旱时间的延长,脯氨酸含量明显增加,在干旱48 h时的提高幅度高于干旱12 h(P<0.05)。在用TSA预处理后再次干旱同样时间,发现脯氨酸含量显著提高了0.57倍和0.34倍,复水后含量明显下降(图4-A)。丙二醛(MDA)随沙棘干旱强度的增加而升高,加速膜脂的过氧化程度。丙二醛含量在干旱12 h后显著上调8.06倍,在干旱48 h后显著上调12.35倍,在复水过程中逐渐下调。而经过TSA预处理后,丙二醛含量在干旱12 h时无显著变化,但在干旱48 h后显著下调(图4-B)。
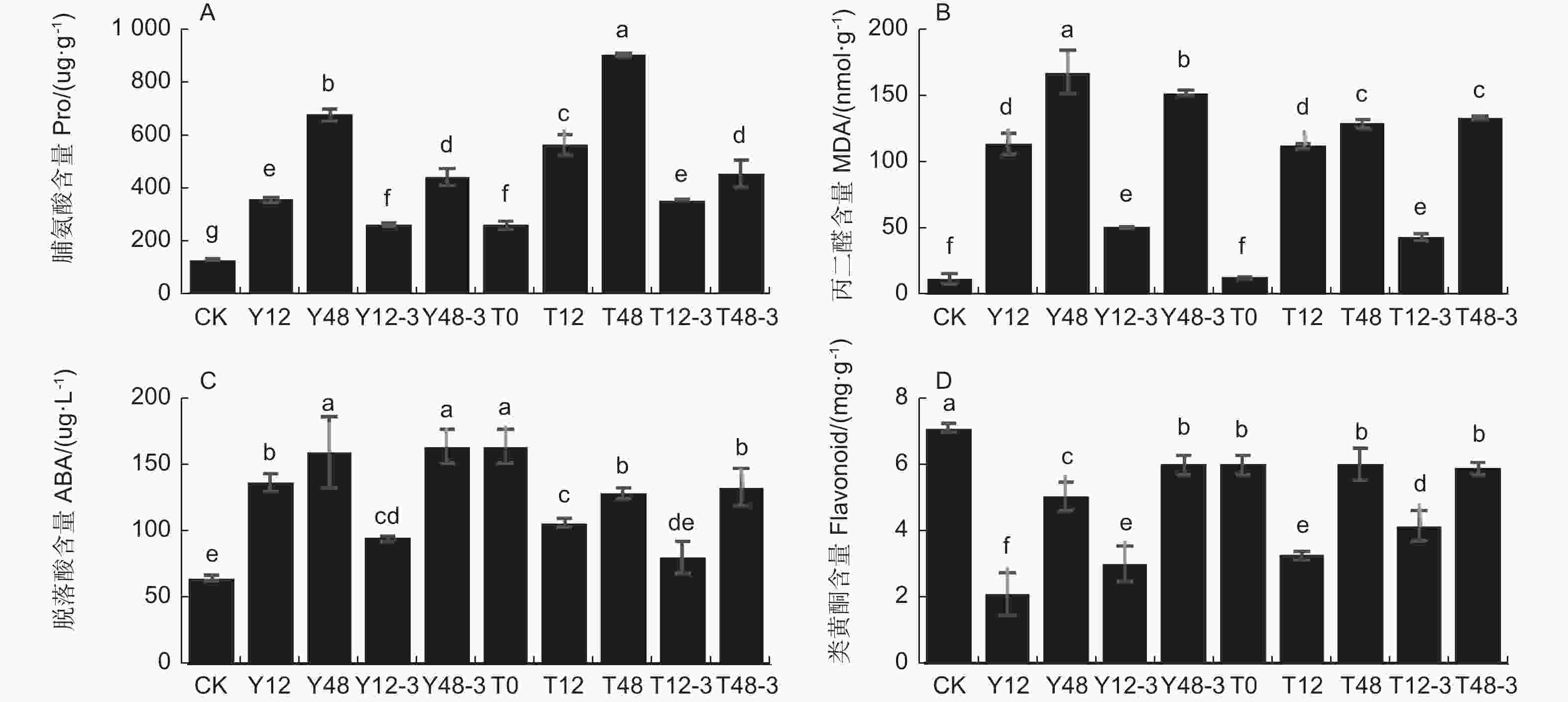
图 4 TSA对沙棘在干旱处理下和复水条件下叶片抗旱生理指标的变化
Figure 4. Effect of TSA on the physiological indexes of sea buckthorn leaf drought resistance under drought treatment and rehydration
干旱处理和TSA处理会影响沙棘叶片脱落酸和黄酮类化合物含量。未使用TSA预处理的干旱12 h条件下,相对于对照组脱落酸含量显著上调,类黄酮含量显著下调, 复水3 d后脱落酸含量显著下调,类黄酮含量显著上调。干旱48 h后相对于对照组脱落酸含量显著上调,类黄酮含量显著下调,复水3 d后脱落酸含量无显著变化,类黄酮含量显著上调(P<0.05)。TSA预处理后的干旱12 h,脱落酸含量相对于对照组显著上调,与仅干旱处理12 h相比增长幅度减少,类黄酮含量相对于对照组显著下调54.4%,与仅干旱处理12 h相比下调幅度减少。复水3 d后脱落酸含量显著下调,类黄酮含量显著上调。TSA预处理后的干旱48 h,脱落酸含量相对于对照组显著上调,与仅干旱处理48 h相比增长幅度减少;类黄酮含量相对于对照组显著下调,与仅干旱处理48 h相比下调幅度减少。复水3 d后脱落酸和类黄酮含量均无显著变化(图4-C, D)。
-
组蛋白去乙酰化酶基因HrHDA6和HrHDA19会影响组蛋白乙酰化程度,从而调控组蛋白乙酰化对抗旱的作用。在未经TSA预处理的干旱条件下,干旱12 h使HrHDA6基因表达量增加3.32倍,HrHDA19-1基因表达量显著增加5.50倍。干旱48 h使HrHDA6基因表达量显著增加6.82倍,HrHDA19-1基因表达量显著增加12.69倍。在TSA预处理后的干旱条件下,干旱12 h使HrHDA6基因表达量显著增加0.48倍,HrHDA19-1基因表达量显著增加1.28倍。干旱48 h使HrHDA6基因表达量显著增加2.25倍,HrHDA19-1基因表达量显著增加2.84倍(P<0.05)。两个基因表达量总体趋势均为干旱处理时最高,TSA预处理后的干旱处理处理次之,对照组最低,且随着干旱时间延长,基因表达量均升高,复水后基因表达量显著降低。在TSA预处理后干旱时间延长,基因表达量均显著升高,干旱12 h后复水基因表达量无显著变化,干旱48 h后复水基因表达量显著降低(图5)。表明沙棘干旱叶片受到TSA影响从而促进了组蛋白去乙酰化酶相关基因的表达。
-
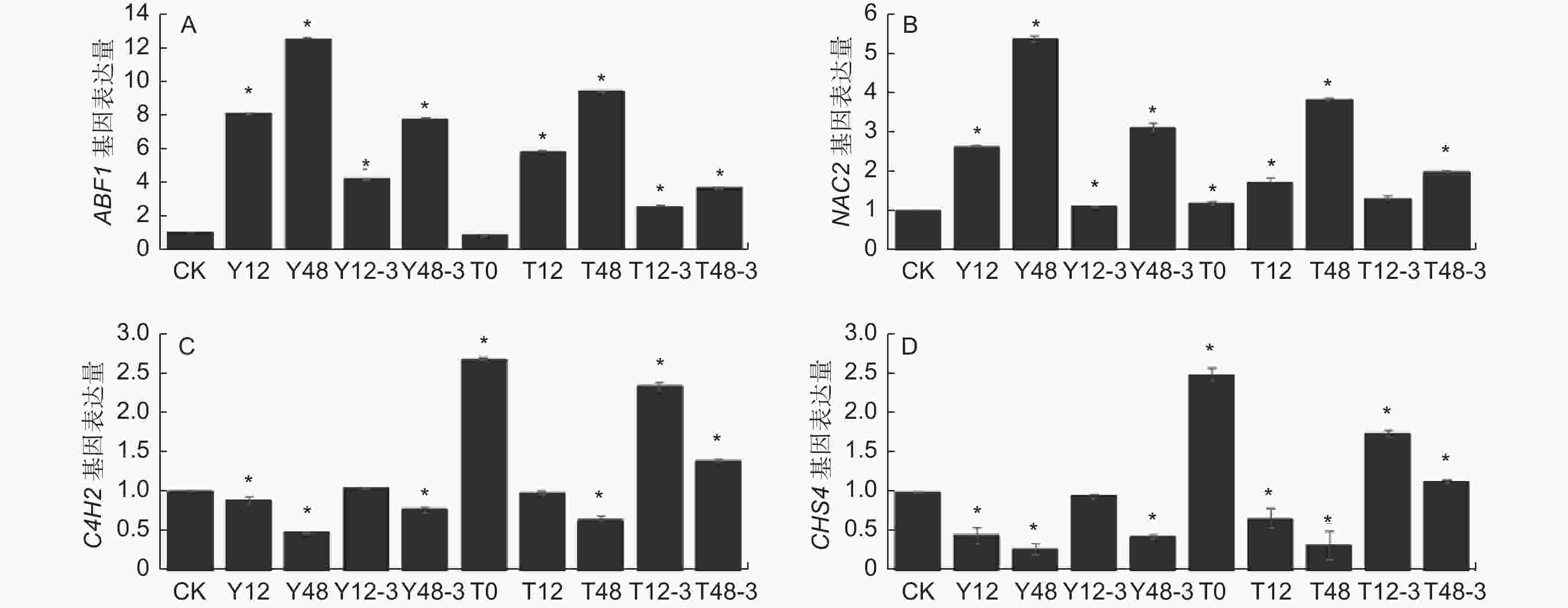
通过前期研究分析发现,沙棘NAC等基因的表达变化会显著影响ABA合成途径,进而影响下游ABF等基因的表达。在未经TSA预处理时,干旱12 h后NAC2基因表达量显著提高1.65倍,ABF1基因表达量显著提高7.09倍。干旱48 h后NAC2基因表达量显著提高5.41倍,ABF1基因表达量显著提高12.57倍。在TSA预处理后的干旱条件下,干旱12 h后NAC2基因表达量轻微上调,ABF1基因表达量显著提高4.83倍,干旱48 h后NAC2基因表达量显著提高2.86倍,ABF1基因表达量显著提高8.45倍(P<0.05)。各个基因表达量总体趋势均为干旱处理时最高,TSA预处理后的干旱处理次之,对照组最低。无论是否经过TSA预处理,随着干旱时间延长,NAC2基因表达量增加幅度均增大,ABF1基因表达量增加幅度均减小,复水后两个基因表达量均显著降低(图6-A, B);肉桂酸4-羟化酶(C4H)和查尔酮合酶(CHS)等基因的表达变化会影响总黄酮合成途径。未经TSA预处理时,干旱12 h和干旱48 h时C4H2 和CHS4基因表达量均下调,且在干旱48 h时下调更大。经过TSA预处理后干旱使C4H2和CHS4基因表达量下调趋势明显减缓(P<0.05)。各个基因表达量总体趋势均为对照组最高,TSA预处理后的干旱时期次之,干旱时期最低。且无论是否经过TSA预处理,随着干旱时间延长,C4H2和CHS4基因表达量均降低,复水后基因表达量均升高(图6-C, D)。表明沙棘叶片组蛋白乙酰化水平上调促进了类黄酮合成相关基因表达,抑制脱落酸合成相关基因的表达。
-
植株处于失水环境会导致其生长缓慢甚至停止生长,茎端和叶片发生下垂及萎蔫现象,影响植物的光合作用、呼吸作用、离子和营养物质的转运以及植物生长调节剂的活性[19],植物抗旱性在表型形态、生理生化和基因表达上与渗透调节和减轻干旱损害的相关指标有关,在气孔运动、光合作用、细胞渗透调节和抗氧化剂等方面会产生明显的表征[20]。沙棘在干旱胁迫下为了提高水分保持能力、减少蒸腾作用和改善光合作用,叶片发生皱缩减少水分蒸腾,植株含水率降低,在干旱程度较低时复水可部分恢复至对照水平,在干旱程度过大时复水无法恢复至对照水平,说明干旱持续时长越长,植株表型受到的损伤越大,越不易恢复生长,而经过TSA预处理的植株在面对同等干旱胁迫时表现出更强的耐旱性。在干旱胁迫处理后,净光合速率、蒸腾速率、气孔导度、Fv/Fm、Y(II)和SPAD值均下调,在经过TSA预处理后下调程度明显减小,说明TSA在一定程度上缓解了干旱胁迫导致沙棘 PSII 的损伤,光合作用速率呈恢复趋势。
植物可通过诱导细胞内的信号级联反应,进而开启或关闭基因表达,从而合成能够对抗压力的所需蛋白质,并合成脯氨酸、丙二醛、脱落酸和类黄酮等物质,以维持细胞内外平衡和保护细胞的结构完整性[21]。植物为了维持细胞的内外膨压,会增加胞内渗透调节分子和细胞防御酶活力,从而有效的清除由于干旱环境导致的有害物质的产生[22]。在干旱胁迫下,沙棘叶片脯氨酸含量明显升高,从而提高渗透保护能力。有研究发现使用25 mg·L−1脯氨酸处理后可以促进干旱胁迫下白榆扦插苗的生长[23]。大豆幼苗在干旱胁迫下拥有着显著变化的脯氨酸含量,干旱胁迫下植株脯氨酸含量是正常浇水植株的5.29倍,而复水后,脯氨酸含量下降到干旱胁迫植株的1/3以下[24],这与沙棘经历干旱和复水阶段后的脯氨酸水平结果类似。MDA含量在干旱胁迫后显著上调8.06~12.35倍,在复水过程中逐渐下调,表明干旱胁迫导致沙棘MDA含量增加,加剧沙棘叶片膜脂过氧化程度,膜选择透性降低,膜完整程度降低,损伤程度呈逐渐增大趋势,这一结果与中国沙棘[25]和小麦(Triticum aestivum L.)[26]相同。
研究发现,大麦(Hordeum vulgare L.)干旱胁迫下4种组蛋白乙酰转移酶(OsHAC703、OsHAG703、OsHAF701和OsHAM701)的表达显著增加[27],而HDA9通过改变拟南芥中逆境响应基因的组蛋白乙酰化水平,从而调节植物对盐和干旱胁迫的敏感性[28],说明干旱胁迫能使组蛋白去乙酰化酶表达升高,而TSA能够抑制沙棘的组蛋白去乙酰化酶,在干旱胁迫下,TSA处理显著降低HrHDA6和HrHDA19-1基因的相对表达量,使沙棘在干旱过程中的组蛋白去乙酰化酶水平降低,进而增强沙棘的抗旱性。
脱落酸等内源植物激素在植株遭受不良环境胁迫时会产生信号响应,从而协调植株能够在干旱胁迫中维持自身生长需要[29-31],脱落酸水平与气孔关闭密切相关[32],植物可以通过关闭叶片上的部分气孔使体内水分的流失程度降低,干旱胁迫下叶片中ABA含量调控气孔运动,提高ABA水平可激活生理反应和信号转导,从而调节植物对脱水的反应和水分利用优化[33]。组蛋白乙酰化已被报道在ABA介导的使植物适应干旱的基因调控中起关键作用[34]。在本研究中发现,未经TSA预处理时,随着干旱程度的加深,ABA合成相关基因NAC2和ABF1表达量逐渐上升,而在TSA预处理后,基因表达量升高趋势减缓,在TSA的作用下脱落酸含量比干旱处理时期增加幅度减少,说明TSA处理加快沙棘叶片气孔关闭, 使其在干旱胁迫下减缓失水的程度, 以缓解干旱造成的影响,保持体内的充足水分。研究发现干旱胁迫下油茶(Camellia oleiferaAbel.)通过增加叶片ABA含量以关闭气孔来减少水分蒸腾,调节自身生长发育和缓解干旱影响[35],ABA反应元件结合蛋白ABP9提高了干旱条件下的光合作用能力[36]。有研究者以大豆幼苗为试验材料,探究干旱胁迫对大豆幼苗生化指标的影响,发现幼苗在经历长期干旱胁迫后,水分亏缺和复水条件下大豆叶片组织内源ABA水平发生了显著变化,ABA含量在第一次干旱胁迫时呈上升趋势,在接下来的干旱胁迫中逐渐下降,复水处理后,植株内源ABA水平降低,与对照植株相似[37]。
而类黄酮可以减少干旱胁迫下氧自由基积累对植物造成的损害[38],TSA预处理后的干旱条件下类黄酮含量相对于仅干旱处理时下调幅度减少,类黄酮合成相关基因C4H2和CHS4的表达量有所升高,说明TSA有效缓解了干旱环境,从而促使氧自由基积累减少,类黄酮合成减少。有研究表明,干旱响应基因SsMYB113通过参与黄酮类化合物和ABA的生物合成,从而增强木荷(Schima superbaGardn. et Champ.)抗旱性[39]。但青钱柳(Cyclocarya paliurus(Batal.)Iljinsk.)在受到不同程度水分胁迫时,黄酮类化合物合成相关的大部分基因会随水分胁迫程度加重而下降[40],这与本研究中干旱48 h后复水3天时类黄酮含量无显著变化结果相同,说明沙棘等植物在长时间干旱胁迫下会使黄酮类化合物合成受阻,无法充分去除干旱造成的氧自由基,从而维持自身生命活动稳定。在课题组前期研究中也发现,干旱胁迫下H3K9乙酰化修饰对6个脱落酸合成和信号通路相关基因进行正调控,对17个类黄酮类生物合成相关基因进行负调控,TSA处理后,干旱胁迫下脱落酸和类黄酮含量及其相关基因表达的变化均有所减缓。
-
本课题研究了TSA对干旱条件下沙棘叶片形态、光合指标、脯氨酸、丙二醛等生理特性和组蛋白去乙酰化酶、脱落酸和类黄酮合成相关基因表达的影响,发现TSA预处理的沙棘在光合生理特性和叶绿素相关指标上的体现的耐旱性更强,TSA通过调节组蛋白去乙酰化酶基因,改变组蛋白乙酰化修饰状态,作用于类黄酮和ABA合成相关基因,从而参与沙棘扦插苗叶片对干旱胁迫和复水过程的响应调节,最终增强沙棘的抗旱性。
曲古抑菌素A对沙棘扦插苗响应干旱和复水及相关基因表达的影响
Effects of Trichostatin A on Responses of Sea Buckthorn to Drought and Rehydration and Related Gene Expression
-
摘要:
目的 研究组蛋白去乙酰化酶抑制剂曲古抑菌素A(Trichostatin A, TSA)在20%聚乙二醇模拟干旱和干旱后复水条件下,对沙棘扦插苗叶片形态、光合指标、脯氨酸、丙二醛和脱落酸含量等生理特性和组蛋白去乙酰化酶、合成脱落酸和类黄酮相关基因表达的影响。 方法 测定沙棘扦插苗干旱相关生理指标,实时定量PCR检测基因表达量。 结果 1 μmol·L−1TSA预处理的沙棘在同等干旱胁迫下耐旱性增强。与仅干旱处理相比,(1)叶片下垂和萎蔫程度降低,植株鲜质量下降程度更小,复水后植株恢复程度更大。(2)净光合速率、蒸腾速率、气孔导度、PSII最大光化学效率Fv/Fm值、PSII有效光化学量子产量Y(Ⅱ)值和叶绿素相对含量(SPAD)值均显著上调,复水后均下调。(3)脯氨酸和类黄酮含量显著上调,丙二醛和脱落酸含量显著下调,复水后趋势相同。(4)组蛋白去乙酰化酶基因HrHDA6和HrHDA19、脱落酸合成相关基因ABF1和NAC2表达均显著下调,类黄酮合成相关基因C4H2和CHS4表达均显著上调,复水后趋势相同。 结论 TSA通过调控沙棘扦插苗生理和基因表达参与对干旱胁迫的响应,可以提高沙棘的抗旱性,该研究为深入解析组蛋白乙酰化影响沙棘抗旱的调控机制奠定重要基础。 Abstract:Objective To study the effects of the histone deacetylase inhibitor Trichostatin A (TSA) on the physiological characteristics of leaf morphology, photosynthetic indexes, proline, malondialdehyde and abscisic acid content, synthetic histone deacetylase, abscisic acid and flavonoid-related genes of sea buckthorn cuttings under the conditions of simulated drought and post-drought rehydration of 20% polyethylene glycol. Methods The drought-related physiological indexes of sea buckthorn cuttings were measured, and the gene expression was detected by quantitative real-time PCR. Result TSA-pretreated (1 μmol·L−1) sea buckthorn was enhanced in drought tolerance under equal drought stress. Compared with the drought treatment, (1) the degree of leaf sagging and wilting reduced, the degree of fresh weight decline was smaller, and the degree of plant recovery after rehydration was greater. (2) The net photosynthetic rate, transpiration rate, stomatal conductivity, PSII maximum photochemical efficiency Fv/Fm value, PSII effective photochemical quantum yield Y(II) value and chlorophyll relative content (SPAD) value all significantly increased, and all were adjusted down after rehydration. (3) The content of proline and flavonoids increased significantly, the content of malondialdehyde and abscisic acid decreased significantly, and the trend was the same after rehydration. (4) The expression of histone deacetylase genes HrHDA6 and HrHDA19, abscisic acid synthesis-related genes ABF1 and NAC2 were significantly down-regulated, and the expression of flavonoid synthesis-related genes C4H2 and CHS4 were significantly upregulated, and the trend after rehydration was the same. Conclusion TSA can improve the drought resistance of sea buckthorn by regulating the physiological and gene expression of sea buckthorn cuttings in response to drought stress, and this study lays an important foundation for in-depth analysis of the regulatory mechanism of histone acetylation affecting sea buckthorn drought resistance. -
表 1 沙棘内参基因和目的基因引物
Table 1. Primers for reference genes and target genes in sea buckthorn
引物名称
Primer name正向引物
Forward primer(5′→3′)反向引物
Reverse primer(5′→3′)功能注释
Functional annotation18S rRNA GCCAAGGAAGTTTGAGGCAA TTCAAAGATTACCCGGGCCT 内参基因 Reference gene ABF2 TCCAGAACACGGTGGGTAGC AAGAGTCAACGAGCCTTGCCT 脱落酸合成途径
Abscisic acid synthesisNAC2 GCTCAAAAGCCGACGACAAG CGAATTCACGCGAGGCAAAT C4H2 AACCCGAAGAGTTCCGACCA CAAGAATAGGCAAAGCCAGAATGAT 类黄酮合成途径
Flavonoid synthesis pathwayCHS4 GTCAACCTAAGTCCAAAATAACCCA CGAAGAACTGTACCTCCAGCAAA HrHDA6 GGGGGGAGCATCTTTACCAT GCTTCATTGGGTGACCTTGG 组蛋白去乙酰化酶基因
Histone deacetylase geneHrHDA19-1 GACCTCCAGACACTGACATTCCA GCTCTCCTGATCTTTAACCTCGG -
[1] 张蓉蓉. 黄土高原矿区典型木本植物幼苗对干旱胁迫的生态适应性研究[D]. 太原: 山西大学, 2015. [2] SHAHID A, NAEEM K, YULIN T. Epigenetic marks for mitigating abiotic stresses in plants[J]. Journal of Plant Physiology, 2022, 275: 153740. doi: 10.1016/j.jplph.2022.153740 [3] BEGCY K, DRESSELHAUS T. Epigenetic responses to abiotic stresses during reproductive development in cereals[J]. Plant Reproduction, 2018, 31(4): 343-355. doi: 10.1007/s00497-018-0343-4 [4] KIM J M, SASAKI T, UEDA M,et al. Chromatin changes in response to drought, salinity, heat, and cold stresses in plants[J]. Frontiers in Plant Science, 2015, 6: 114. [5] DU Q L, FANG Y P, JIANG J M,et al. Characterization of histone deacetylases and their roles in response to abiotic and PAMPs stresses in Sorghum bicolor[J]. BMC Genomics, 2022, 23(1): 28. doi: 10.1186/s12864-021-08229-2 [6] ZHANG J B, HE S P, LUO J W,et al. A histone deacetylase, GhHDT4D, is positively involved in cotton response to drought stress[J]. Plant Molecular Biology, 2020, 104(1-2): 67-79. doi: 10.1007/s11103-020-01024-9 [7] SHEN Y, LEI T T, CUI X Y,et al. Arabidopsis histone deacetylase HDA15 directly represses plant response to elevated ambient temperature[J]. The Plant Journal:for Cell and Molecular Biology, 2019, 100(5): 991-1006. doi: 10.1111/tpj.14492 [8] YANG C, SHEN W J, CHEN H F,et al. Characterization and subcellular localization of histone deacetylases and their roles in response to abiotic stresses in soybean[J]. BMC Plant Biology, 2018, 18(1): 226. doi: 10.1186/s12870-018-1454-7 [9] GUO J E, WANG H H, YANG Y,et al. Histone deacetylase gene SlHDA3 is involved in drought and salt response in tomato[J]. Plant Growth Regulation, 2023, 99(2): 359-372. doi: 10.1007/s10725-022-00913-x [10] 陈勇智, 苏良辰, 李 玲. TSA对不同抗旱性花生品种响应干旱过程中光合特性及相关基因表达的影响[J]. 华南师范大学学报(自然科学版), 2018, 50(3):72-77. [11] ZHANG B H, SU L C, HU B,et al. Expression of AhDREB1, an AP2/ERF transcription factor gene from peanut, is affected by histone acetylation and increases abscisic acid sensitivity and tolerance to osmotic stress in Arabidopsis[J]. International Journal of Molecular Sciences, 2018, 19(5): 1441. doi: 10.3390/ijms19051441 [12] XING, LIU,. AhHDA1-mediated AhGLK1 promoted chlorophyll synthesis and photosynthesis regulates recovery growth of peanut leaves after water stress[J]. Plant Science, 2020, 294: 110461. doi: 10.1016/j.plantsci.2020.110461 [13] HU Y, ZHANG L, ZHAO L,et al. Trichostatin A selectively suppresses the cold-induced transcription of the ZmDREB1 gene in maize[J]. PLoS ONE, 2011, 6(7): e22132. doi: 10.1371/journal.pone.0022132 [14] SONG Y, LIU L J, LI G P,et al. Trichostatin A and 5-Aza-2'-Deoxycytidine influence the expression of cold-induced genes in Arabidopsis[J]. Plant Signal Behav, 2017, 12(11): e1389828. doi: 10.1080/15592324.2017.1389828 [15] GAO G R, LV Z R, ZHANG G Y,et al. An ABA-flavonoid relationship contributes to the differences in drought resistance between different sea buckthorn subspecies[J]. Tree Physiology, 2021, 41(5): 744-755. doi: 10.1093/treephys/tpaa155 [16] ZHANG T, GAO G R, LIU J J,et al. Transcripts and ABA-dependent signaling in response to drought stress in Hippophae rhamnoides L[J]. Trees, 2020, 34(4): 1033-1045. doi: 10.1007/s00468-020-01979-8 [17] LIVAK K J, SCHMITTGEN T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method[J]. Methods, 2001, 25(4): 402-408. doi: 10.1006/meth.2001.1262 [18] 王 谦, 陈泠澍, 赵 薇, 等. 干旱条件下栓皮栎叶片光系统活性受损的温度指标[J]. 中国农业气象, 2019, 40(5):308-316. [19] FAROOQ M, WAHID A, KOBAYASHI N, et al. Plant drought stress: Effects, mechanisms and management[M]//Sustainable Agriculture. Dordrecht: Springer Netherlands, 2009: 153-188. [20] Muchate N S, Nikalje G C, Rajurkar N S,et al. Plant salt stress: adaptive responses, tolerance mechanism and bioengineering for salt tolerance[J]. Botanical Review, 2016, 82(4): 371-406. doi: 10.1007/s12229-016-9173-y [21] BANERJEE A, ROYCHOUDHURY A. Group II late embryogenesis abundant (LEA) proteins: Structural and functional aspects in plant abiotic stress[J]. Plant Growth Regulation, 2016, 79(1): 1-17. doi: 10.1007/s10725-015-0113-3 [22] Fang Y and Xiong L. General mechanisms of drought response and their application in drought resistance improvement in plants[J]. Cellular and Molecular Life Sciences, 2015, 72(4): 673-689. doi: 10.1007/s00018-014-1767-0 [23] 张 晓, 曹慧芬, 赵建国, 等. 脯氨酸对干旱胁迫下白榆扦插苗生长的影响[J]. 山西大同大学学报(自然科学版), 2021, 37(3):1-4. [24] Nguyen T, Le T, Pham H,et al. Evaluation of proline, soluble sugar and ABA content in soybean Glycine max (L. ) under drought stress memory[J]. AIMS Bioengineering, 2020, 7(3): 114-123. doi: 10.3934/bioeng.2020011 [25] 韩蕊莲, 李丽霞, 梁宗锁. 干旱胁迫下沙棘叶片细胞膜透性与渗透调节物质研究[J]. 西北植物学报, 2003, 23(1):23-27. doi: 10.3321/j.issn:1000-4025.2003.01.005 [26] 白志英, 李存东, 吴同燕, 等. 干旱胁迫条件下小麦旗叶酶活性和丙二醛含量的染色体定位[J]. 植物遗传资源学报, 2009, 10(2):255-261. doi: 10.13430/j.cnki.jpgr.2009.02.014 [27] DEMETRIOU K, KAPAZOGLOU A, TONDELLI A,et al. Epigenetic chromatin modifiers in barley: I. Cloning, mapping and expression analysis of the plant specific HD2 family of histone deacetylases from barley, during seed development and after hormonal treatment[J]. Physiologia Plantarum, 2009, 136(3): 358-368. doi: 10.1111/j.1399-3054.2009.01236.x [28] ZHENG Y, DING Y, SUN X,et al. Histone deacetylase HDA9 negatively regulates salt and drought stress responsiveness in Arabidopsis[J]. Journal of Experimental Botany, 2016, 67(6): 1703-1713. doi: 10.1093/jxb/erv562 [29] Bano A, Hansen H, DÖRFFLING K,et al. Changes in the contents of free and conjugated abscisic acid, phaseic acid and cytokinins in xylem sap of drought stressed sunflower plants[J]. Phytochemistry, 1994, 37(2): 345-347. doi: 10.1016/0031-9422(94)85058-5 [30] Peleg Z, Blumwald E. Hormone balance and abiotic stress tolerance in crop plants[J]. Current Opinion in Plant Biology, 2011, 14(3): 290-295. doi: 10.1016/j.pbi.2011.02.001 [31] Zhang J H, Jia W S, Yang J C,et al. Role of ABA in integrating plant responses to drought and salt stresses[J]. Field Crops Research, 2006, 97(1): 111-119. doi: 10.1016/j.fcr.2005.08.018 [32] Sharma A, Shahzad B, Kumar V,et al. Phytohormones regulate accumulation of osmolytes under abiotic stress[J]. Biomolecules, 2019, 9(7): 285-321. doi: 10.3390/biom9070285 [33] Song L, Huang S, Wise A,et al. A transcription factor hierarchy defines an environmental stress response network[J]. Science, 2016, 354(6312): 1550. doi: 10.1126/science.aag1550 [34] SRIDHA S, WU K Q. Identification of AtHD2C as a novel regulator of abscisic acid responses in Arabidopsis[J]. The Plant Journal:for Cell and Molecular Biology, 2006, 46(1): 124-133. doi: 10.1111/j.1365-313X.2006.02678.x [35] HE X S, XU L C, PAN C,et al. Drought resistance of Camellia oleifera under drought stress: Changes in physiology and growth characteristics[J]. PLoS ONE, 2020, 15(7): e0235795. doi: 10.1371/journal.pone.0235795 [36] ZHANG X, WOLLENWEBER B, JIANG D,et al. Water deficits and heat shock effects on photosynthesis of a transgenic Arabidopsis thaliana constitutively expressing ABP9, a bZIP transcription factor[J]. Journal of Experimental Botany, 2008, 59(4): 839-848. doi: 10.1093/jxb/erm364 [37] GUPTA A, RICOMEDINA A, CAÑODELGADO A I. The physiology of plant responses to drought[J]. Science, 2020, 368(6488): 266-269. doi: 10.1126/science.aaz7614 [38] MA D Y, SUN D X, WANG C Y,et al. Expression of flavonoid biosynthesis genes and accumulation of flavonoid in wheat leaves in response to drought stress[J]. Plant Physiology and Biochemistry, 2014, 80: 60-66. doi: 10.1016/j.plaphy.2014.03.024 [39] ZHANG G H, YU Z M, YAO B,et al. SsMYB113, a Schima superba MYB transcription factor, regulates the accumulation of flavonoids and functions in drought stress tolerance by modulating ROS generation[J]. Plant and Soil, 2022, 478(1-2): 427-444. doi: 10.1007/s11104-022-05466-6 [40] SHENG X L, CHEN H W, WANG J M,et al. Joint transcriptomic and metabolic analysis of flavonoids in Cyclocarya paliurus leaves[J]. ACS Omega, 2021, 6(13): 9028-9038. doi: 10.1021/acsomega.1c00059 -



 下载:
下载: