-
bHLH(basic Helix-Loop-Helix, 碱性螺旋-环-螺旋)转录因子是植物中广泛存在的一类大的转录因子家族,其结构域由约60个氨基酸组成,包含N端碱性区域和C端螺旋-环-螺旋区域[1-3]。目前研究发现,bHLH转录因子主要通过识别E-box(CANNTG)来调控下游靶基因,E-box的形式多种多样,其中, G-box(CACGTG)最常见[4-5]。已有研究表明,bHLH转录因子在植物生长发育、逆境反应、信号转导等一系列过程中起重要作用[1, 6], 如,bHLHs参与雌蕊发育[7]、花期控制[8]、根毛的分化[9]、花器官和叶腋分生组织形成[10]、光信号途径[11]、油菜素内酯和脱落酸信号传导[12]。其中,bHLH对抗逆的调控作用尤其值得关注,目前已经发现和鉴定了多种环境胁迫应答的bHLH转录因子,并发现bHLH转录因子在植物抗逆方面的调控作用, 如,拟南芥bHLH122能够调控气孔的开度,抑制水分散失速率,同时抑制ABA的分解,能够调控拟南芥的抗旱、耐盐能力[2]; 拟南芥bHLH92转录因子参与调控干旱和渗透胁迫响应[13]; 拟南芥ICE1基因正向调控冷害响应过程[14]。bHLH蛋白(JAM1)可负调控茉莉酸(JA)信号途径,在JA介导的植物应激反应中起到举足轻重的作用[15]。水稻OsbHLH148基因在抗旱过程中作为一个起始应答因子调节受JA调节的基因的表达[16]。拟南芥中过表达OrbHLH2或OrbHLH001基因提高了拟南芥的耐盐、抗冷和渗透胁迫能力[17]。小麦TabHLH1基因主要通过ABA依赖途径在植物的抗渗透胁迫中起到至关重要的作用[18]。上述研究表明,bHLH转录因子在调控植物抗逆过程中起重要作用。
刚毛柽柳(Tamarix hispida Willd.)属柽柳科柽柳属植物,是盐生木本植物,灌木或小乔木。柽柳具有很强的耐盐碱、干旱和高温能力,是研究木本植物抗逆分子机理,分离重要耐盐功能基因的理想物种之一。前期研究工作表明,刚毛柽柳ThbHLH1基因(GenBank登录号: KM101094)能够响应盐、干旱等非生物胁迫,且在根、茎及叶器官的表达模式不同;过表达ThbHLH1能够提高柽柳的耐盐、抗旱能力;利用酵母单杂交技术,鉴定出ThbHLH1转录因子能够识别G-box元件,并且在盐或干旱胁迫下,ThbHLH1通过结合G-box来激活基因表达的能力显著增强[19]。为了进一步研究ThbHLH1转录因子还可通过识别哪些元件来调控基因的表达,笔者我们利用以转录因子为中心的酵母单杂交技术(TF-centered Y1H)[20],通过对随机DNA文库的筛选,来鉴定ThbHLH1转录因子识别的新元件。本研究将为鉴定ThbHLH1转录因子直接调控的下游靶基因奠定基础。
-
刚毛柽柳种植于沙与草炭土混合物中(比例为3:1),取2月苗龄植株用于实验研究。小叶烟草(Nicotiana tabacum L.)种植于草炭土:蛭石:珍珠岩混合物中(比例为3:1:1),取6周苗龄植株用于实验研究。植物材料置于温度(22±2)℃、相对湿度65%75%、光照强度400 μmol·m-2·s-1、光周期16 h光照/8 h黑暗的人工气候室培养。
-
在ThbHLH1基因的5’端和3’端分别引入Sma I线性化的pGADT7-Rec2载体的同源序列,设计引物(表 1)。以柽柳cDNA为模板,扩增出含有pGADT7-Rec2载体同源序列的ThbHLH1基因,将其用同源融合方法连接到pGADT7-Rec2载体上,构建pGADT7-Rec2-ThbHLH1酵母表达载体。
表 1 构建pGADT7-Rec2-ThbHLH1载体引物序列
Table 1. Primer sequences used in construction of the recombinant vector pGADT7-Rec2-ThbHLH1
引物名称
Primer name引物序列(5’-3’)
Sequence (5’-3’)Rec2-ThbHLH1 F AAGCAGTGGTATCAACGCAGAGTGGCCATTATGGCCCATGGCTAATAATCCAGGAG Rec2-ThbHLH1 R TCTAGAGGCCGAGGCGGCCGACATGTTATGAAAGGGGATTTGTC 注:_为pGADT7-Rec2载体的同源序列。 Note:_represents the homologous sequences of pGADT7-Rec2 vector. -
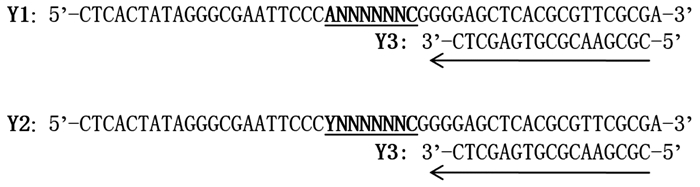
根据酵母单杂交系统(BD Matchmaker TM One-Hybrid Library Construction & Screening Kit. Clontech, Palo Alta, CA, USA)中酵母报告载体pHIS2的序列,设计Y1和Y2两条简并引物,一条特异性引物Y3(表 2),分别以Y1和Y2为模板,Y3为引物进行PCR扩增(图 1)。将扩增产物连接到pHIS2载体上,构建随机DNA文库,具体方法见参考文献[20]。
表 2 构建随机DNA文库引物序列
Table 2. Primers used in constructing a random short DNA sequence insertion library
引物名称
Primer name引物序列(5’-3’)
Sequence (5’-3’)Y1 
Y2 
Y3 CGCGAACGCGTGAGCTC 注:_为Sma I线性化pHIS2载体两端的序列, 为随机DNA序列, N: A/G/C/T;Y: C/T Note:_represents the sequences on both ends of the Sma I linearized pHIS2 vector, represents the random DNA sequences, N: A/G/C/T;Y: C/T -
按照酵母单杂交系统操作说明书进行Y187酵母感受态细胞的制备。将效应载体pGADT7-Rec2-ThbHLH1和随机DNA文库共转化Y187酵母感受态细胞,筛选柽柳ThbHLH1转录因子识别的顺式作用元件。具体方法如下:向500 μL Y187酵母感受态细胞中加入2 μg效应载体pGADT7-Rec2-ThbHLH1和1.5 μg随机DNA文库质粒,共转化Y187酵母感受态细胞。将转化后的产物分别涂布于SD/-Trp、SD/-Leu、SD/-Leu/-Trp(DDO)和SD/-His/-Leu/-Trp(TDO)+30 mmol·L-1 3-AT的筛选培养基上,30℃倒置培养3~5 d。
-
挑取TDO+30 mmol·L-1 3-AT筛选培养基上的阳性克隆菌落,于TDO液体培养基中30℃震荡(250 r·min-1)培养12 d;提取酵母质粒,转化大肠杆菌,提取质粒进行测序。测序结果与pHIS2载体序列进行比对,获得pHIS2载体插入的DNA序列,然后通过PLACE(http://www.dna.affrc.go.jp/PLACE/)和PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/)分析插入序列中是否含有已知的顺式作用元件。
将顺式作用元件及其突变元件的寡核苷酸片段3次串联,构建到pHIS2载体上,引物见表 3。将构建好的效应载体(pGADT7-Rec2-ThbHLH1)和重组报告载体(pHIS2-Cis)共转化Y187酵母感受态细胞,转化后的酵母分别涂布于DDO和TDO+30 mmol·L-1 3-AT固体培养基上,30℃倒置培养35 d。挑取阳性克隆菌落,于TDO液体培养基中30℃震荡培养1~2 d,将菌液分别稀释10、100、1 000、10 000倍,取2 μL点于TDO+30 mmol·L-1 3-AT培养基上,30℃倒置培养2~3 d,拍照。
表 3 构建酵母报告载体pHIS2-Cis引物序列
Table 3. Primer sequences used in constructing pHIS2-Cis yeast reporter vectors
引物名称
Primer name引物序列(5’-3’)
Sequence (5’-3’)pHIS2-LTRE1F AATTCCCGAAACCGAAACCGAAAGAGCT pHIS2-LTRE1R CTTTCGGTTTCGGTTTCGGG pHIS2-L1M1F AATTCAAGAAAAAGAAAAAGAAAGAGCT pHIS2-L1M1R CTTTCTTTTTCTTTTTCTTG pHIS2-L1M2F AATTCCCAAAACCAAAACCAAAAGAGCT pHIS2-L1M2R CTTTTGGTTTTGGTTTTGGG pHIS2-L1M3F AATTCCCGCCACCGCCACCGCCAGAGCT pHIS2-L1M3R CTGGCGGTGGCGGTGGCGGG pHIS2-L1M4F AATTCCCGAACCCGAACCCGAACGAGCT pHIS2-L1M4R CGTTCGGGTTCGGGTTCGGG pHIS2-L1M5F AATTCAAAAAAAAAAAAAAAAAAGAGCT pHIS2-L1M5R CTTTTTTTTTTTTTTTTTTG pHIS2-L1M6F AATTCCCGCCCCCGCCCCCGCCCGAGCT pHIS2-L1M6R CGGGCGGGGGCGGGGGCGGG pHIS2-L1M7F AATTCAAACCCAAACCCAAACCCGAGCT pHIS2-L1M7R CGGGTTTGGGTTTGGGTTTG pHIS2-WRKY710SF AATTCTGACTGACTGACGAGCT pHIS2-WRKY710SR CGTCAGTCAGTCAG pHIS2-WM1F AATTCCGACCGACCGACGAGCT pHIS2-WM1R CGTCGGTCGGTCGG pHIS2-WM2F AATTCTAACTAACTAACGAGCT pHIS2-WM2R CGTTAGTTAGTTAG pHIS2-WM3F AATTCTGCCTGCCTGCCGAGCT pHIS2-WM3R CGGCAGGCAGGCAG pHIS2-WM4F AATTCTGAATGAATGAAGAGCT pHIS2-WM4R CTTCATTCATTCAG pHIS2-WM5F AATTCCAACCAACCAACGAGCT pHIS2-WM5R CGTTGGTTGGTTGG pHIS2-WM6F AATTCTGCATGCATGCAGAGCT pHIS2-WM6R CTGCATGCATGCAG pHIS2-WM7F AATTCCACACACACACAGAGCT pHIS2-WM7R CTGTGTGTGTGTGG 注:_为粘性末端。Note: represents the sticky ends. -
将顺式作用元件及其突变元件的寡核苷酸片段3次串联,与CaMV 35S minimal promoter(46 bp)连接,引入Hind Ⅲ和NcoⅠ酶切位点,进行DNA序列合成。将合成的序列定向连接到pCAMBIA1301载体上,分别构建报告载体pCAM-LTRE1、pCAM-L1M7、pCAM-WRKY710S和pCAM-WM7(图 2A,引物见表 4),pCAM-G-box、pCAM-GM7载体为本实验室构建保存,上述报告载体统称为pCAM-Cis。将ThbHLH1基因定向连接到pROKⅡ表达载体上,构建效应载体pROKⅡ-ThbHLH1(图 2A,引物见表 4)。
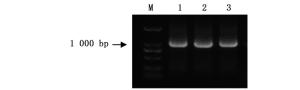
图 2 盐、干旱胁迫下ThbHLH1对G-box、LTRE1及WRKY710S元件的识别能力
Figure 2. The binding affinity of ThbHLH1 to G-box, LTRE1 or WRKY710S in response to salt or drought stress
表 4 构建报告载体及效应载体引物序列
Table 4. Primer sequences used in constructing reporter and effector vectors
引物名称
Primer name引物序列(5’-3’)
Sequence (5’-3’)pCAM-LTRE1F AGCTTCCGAAACCGAAACCGAAAACCCTTCCTCTATATAAGGAAGTTCATTTCATTTGGAGAGAACACGGC pCAM-LTRE1R CATGGCCGTGTTCTCTCCAAATGAAATGAACTTCCTTATATAGAGGAAGGGTTTTCGGTTTCGGTTTCGGA pCAM-L1M7F AGCTTAAACCCAAACCCAAACCCACCCTTCCTCTATATAAGGAAGTTCATTTCATTTGGAGAGAACACGGC pCAM-L1M7R CATGGCCGTGTTCTCTCCAAATGAAATGAACTTCCTTATATAGAGGAAGGGTGGGTTTGGGTTTGGGTTTA pCAM-WRKY710SF AGCTTTGACTGACTGACACCCTTCCTCTATATAAGGAAGTTCATTTCATTTGGAGAGAACACGGC pCAM-WRKY710SR CATGGCCGTGTTCTCTCCAAATGAAATGAACTTCCTTATATAGAGGAAGGGTGTCAGTCAGTCAA pCAM-WM7F AGCTTCACACACACACAACCCTTCCTCTATATAAGGAAGTTCATTTCATTTGGAGAGAACACGGC pCAM-WM7R CATGGCCGTGTTCTCTCCAAATGAAATGAACTTCCTTATATAGAGGAAGGGTTGTGTGTGTGTGA pROKⅡ-ThbHLH1 F 
pROKⅡ-ThbHLH1 R 
注:_为粘性末端, 
Note:_ represents the sticky ends, 
-
选取约2 cm×2 cm大小的烟草叶片,背面向上平铺于1/2 MS培养基上,暗培养2 d,备用。将构建的报告载体(pCAM-Cis)、效应载体(35S:ThbHLH1)及35S:LUC质粒混合(2:2:1),按照PDS-1000基因枪操作法瞬时转化烟草叶片,将转化后的烟草置于人工气候室中暗培养48 h,分别测定GUS和LUC酶活,计算GUS/LUC比值,方法参见Jefferson等[21]。
-
ThbHLH1基因用同源融合的方法连接到pGADT7-Rec2载体上。菌落PCR结果显示:扩增条带特异且大小与预期一致(图 3)。用pGADT7-Rec2F/R载体引物进行测序,确定pGADT7-Rec2-ThbHLH1酵母效应载体构建成功。
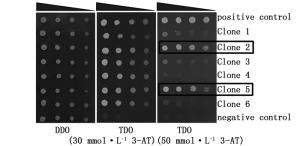

图 3 pGADT7-Rec2-ThbHLH1载体构建菌落PCR检测
Figure 3. PCR analysis of the Escherichia coli containing pGADT7-Rec2-ThbHLH1 vector
根据酵母报告载体pHIS2序列设计的Y1和Y2引物中包含7碱基序列:“ANNNNNN”、“TNNNNNN”、“CNNNNNN”和“GNNNNNN”(反补序列为Y2中的“NNNNNNC”);也包含8碱基序列:“ANNNNNNC”、“CNNNNNNC”和“TNNNNNNC”。因此,理论上利用酵母单杂交系统筛选上述随机DNA文库,能鉴定出ThbHLH1转录因子所识别的不超过7或8个碱基的任意DNA序列。将PCR产物连接到pHIS2载体上,收集所有阳性克隆提取质粒,即为随机DNA文库。
-
为了鉴定ThbHLH1转录因子识别的DNA序列,将酵母效应表达载体(pGADT7-Rec2-ThbHLH1)与随机DNA文库质粒共转化酵母,通过酵母单杂交系统进行筛选,在TDO+ 30 mmol·L-1 3-AT培养基上共获得6个阳性克隆菌落,经TDO+50 mmol·L-1 3-AT培养基进一步筛选后,有2个克隆可正常生长,且长势与阳性对照(p53HIS2/pGADT7-Rec2-p53)的长势一致,即克隆2和5;其余4个克隆则无法正常生长(图 4)。因此,笔者进一步研究这2个阳性克隆。通过酵母质粒提取试剂盒提取酵母质粒,将其转化大肠杆菌感受态细胞,选取阳性克隆进行测序,将测序结果与pHIS2载体序列进行比对,获得2个插入DNA序列。通过PLACE和PlantCARE对获得序列进行顺式作用元件预测,来分析ThbHLH1转录因子所识别的顺式作用元件,结果见表 5,其中,“CCCCCCGAAACGGG”包含LTRE1元件(CCGAAA);“CCCCGCTGACCGGG”包含WRKY710S元件(TGAC)。
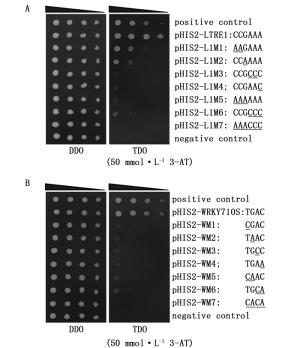
图 4 ThbHLH1转录因子筛选随机DNA文库结果
Figure 4. Interaction of ThbHLH1 with the random DNA sequence insertion library.
表 5 ThbHLH1转录因子识别的插入序列分析
Table 5. Analysis of the insertion sequences bind to ThbHLH1
随机DNA插入序列
(5′-3′)Random DNA insertion
sequences (5′-3′)元件预测
Motif prediction参考文献
ReferencesCCCCCCGAAACGGG LTRE1: “CCGAAA” [22] CCCCGCTGACCGGG WRKY710S: “TGAC” [23] 注:_为文库的插入序列,左侧的CCC及右侧的GGG为载体侧翼序列,因为载体序列也可能和插入序列形成新的元件,所以要考虑左右两端各3个碱基的插入序列。 Note: _ represents the insertion sequences of the library, the left CCC and the right GGG are the vector flanking sequences, the insertion sequences of the three bases on the left and right sides should be considered, because the vector sequence may also form new motifs with the insertion sequences. -
将效应载体(pGADT7-Rec2-ThbHLH1)分别和pHIS2-LTRE1、pHIS2-WRKY710S及其突变载体(pHIS2-L1M1-7、pHIS2-WM1-7)共转化酵母,通过DDO和TDO+30 mmol·L-1 3-AT固体培养基30℃倒置培养3~5 d。挑取DDO或TDO+30 mmol·L-1 3-AT上生长的酵母菌落接种于DDO或TDO液体培养基中,30℃震荡培养至OD600=0.81.0,菌液适当稀释后取2 μL,分别点于DDO和TDO+50 mmol·L-1 3-AT固体筛选培养基上,30℃倒置培养23 d,结果(图 5A、B)表明:pGADT7-Rec2-ThbHLH1分别与pHIS2-LTRE1或pHIS2-WRKY710S共转化的酵母在含有50 mmol·L-1 3-AT的TDO筛选培养基上可正常生长,且长势与阳性对照(p53HIS2/pGADT7-Rec2-p53)的长势一致。LTRE1元件不同碱基位点突变的报告载体(pHIS2-L1M1-7)与pGADT7-Rec2-ThbHLH1共转化的酵母在含50 mmol·L-1 3-AT的TDO筛选培养基上均长势较弱或无法正常生长;而WRKY710S元件不同碱基位点突变的报告载体(pHIS2-WM1-7)与pGADT7-Rec2-ThbHLH1共转化的酵母在含50 mmol·L-1 3-AT的TDO筛选培养基上均无法正常生长。上述结果表明:ThbHLH1转录因子可识别LTRE1和WRKY710S顺式作用元件。
-
为了验证在植物体内ThbHLH1转录因子对LTRE1和WRKY710S元件的识别能力,通过基因枪法将报告载体(pCAM-Cis)、效应载体(35S:ThbHLH1)及35S-LUC质粒共转化烟草叶片,通过测定GUS和LUC酶活性,计算GUS/LUC比值来均一化遗传转化效率,结果见图 2B。效应载体(35S:ThbHLH1)分别与报告载体pCAM-G-box、-LTRE1及-WRKY710S共转化烟草叶片,其GUS/LUC的比值较高,且在盐和干旱胁迫条件下,GUS/LUC比值显著增加,与阳性对照值(单独转化pCAMBIA1301)相比,说明GUS基因大量表达;而pROKII-ThbHLH1分别与上述元件的突变元件共转化烟草,则GUS/LUC值较低,且在盐和干旱胁迫条件下GUS/LUC值无明显变化,与阴性对照值(单独转化报告载体pCAM-Cis)基本一致,说明GUS基因未表达。上述结果表明:在盐和干旱胁迫条件下,ThbHLH1能够识别G-box、LTRE1和WRKY710S元件来激活基因的表达,而不能与其突变元件结合来激活基因的表达(图 2B),该结果与酵母单杂交结果一致,说明在植物体内ThbHLH1转录因子除识别G-box元件外,还可识别LTRE1和WRKY710S元件。
-
转录因子在响应由环境因素引起的基因表达调控方面发挥着重要的作用[24]。它能够通过特异的DNA-蛋白质、蛋白质-蛋白质之间的相互作用,调节下游靶基因的表达,从而抵御外界环境胁迫[25-26]。bHLH转录因子通过识别G-box元件参与植物非生物胁迫的应答,在植物的抗逆过程中起重要作用[4-5],而对bHLH转录因子识别LTRE1和WRKY710S元件参与植物非生物胁迫的应答未见报道。在前期的研究工作中,笔者鉴定出柽柳ThbHLH1转录因子可识别G-box元件,并且在盐或干旱胁迫下,ThbHLH1通过结合G-box元件来激活基因表达的能力显著增强[19]。本研究通过TF-Centered Y1H技术[20]鉴定出ThbHLH1转录因子可识别LTRE1和WRKY710S顺式作用元件(图 5),为了验证在植物体内ThbHLH1转录因子能够识别上述元件,本研究利用共转化reporter和effector法对烟草叶片进行瞬时转化。在盐或干旱胁迫条件下,进行GUS酶活测定及分析,发现ThbHLH1与G-box元件结合后对基因的转录激活能力最强,与LTRE1元件结合后对基因的转录激活能力最弱(图 2B)。这些数据表明,在植物体内ThbHLH1转录因子可通过识别G-box、WRKY710S及LTRE1顺式作用元件来激活基因表达。
Dunn等[22]鉴定出大麦(Hordeum vulgare L.)低温响应基因(blt4.9)启动子中的LTRE1(CCGAAA)元件,该元件参与大麦blt4.9基因的低温应答。Xue[27]研究发现,HvCBF1转录因子通过结合LTRE15(CCGAC)和LTRE1(CCGAAA)元件来调控大麦中的冷应答基因。Matton等[23]鉴定出马铃薯(Solanum tuberosum L.)病程相关基因STH-2启动子中含有WRKY710S (TGAC)元件,该元件参与马铃薯STH-2基因的调控。本研究鉴定出ThbHLH1转录因子除识别G-box元件外,还可识别LTRE1和WRKY710S元件。目前,大量研究表明,bHLH转录因子通过识别相应的顺式作用元件来参与植物耐寒途径的调控,如Feng等[28]从苹果中分离鉴定的一个低温响应基因MdCIbHLH1,该基因能特异结合到拟南芥CBFs启动子的MYC识别位点并激活其表达。Huang等[29]从枳PtrbHLH中克隆到受低温诱导的基因,发现PtrbHLH可以结合POD基因启动子区域中的E-box元件,通过调控POD介导的H2O2清除参与植物对冷胁迫信号的应答,来提高过表达烟草或柠檬的耐寒性。鉴于上述工作,我们推测柽柳ThbHLH1转录因子不仅通过识别G-box元件参与植物耐盐、抗旱途径调控,还可能通过与LTRE1或WRKY710S元件结合来调控耐寒、抗病等相关途径。在后续试验中,笔者将开展ThbHLH1转录因子耐寒、抗病等相关研究工作,同时进行逆境胁迫下过表达及野生型柽柳的转录组测序,找出ThbHLH1转录因子调控的下游靶基因。通过分析靶基因启动子中是否含有G-box、LTRE1及WRKY710顺式作用元件,来鉴定ThbHLH1转录因子直接调控的下游靶基因。
-
本研究通过TF-Centered Y1H技术鉴定出刚毛柽柳ThbHLH1转录因子除识别G-box元件外,还可识别LTRE1及WRKY710S顺式作用元件。利用共转化reporter和effector法对烟草叶片进行瞬时转化,在盐或干旱胁迫条件下,进行GUS酶活测定及分析,发现ThbHLH1与G-box元件结合后对基因的转录激活能力最强,与LTRE1元件结合后对基因的转录激活能力最弱。结果表明,在植物体内ThbHLH1转录因子可通过识别G-box、WRKY710S及LTRE1顺式作用元件来激活基因表达。本研究为研究ThbHLH1转录因子抗逆功能提供了新思路,为进一步揭示柽柳ThbHLH1转录因子抗逆分子调控机制奠定了基础。
刚毛柽柳ThbHLH1转录因子识别的顺式作用元件的鉴定
Identification of Cis-acting Elements Recognized by ThbHLH1 Transcription Factor of Tamarix hispida
-
摘要:
目的 本研究拟对ThbHLH1所识别的顺式作用元件进行鉴定,进一步揭示ThbHLH1调控抗逆基因表达的机理。 方法 利用以转录因子为中心的酵母单杂交系统鉴定ThbHLH1所识别的顺式作用元件;将元件与报告基因融合构建报告载体(pCAM-Cis),通过基因枪法将报告载体与效应载体35S:ThbHLH1共转化烟草叶片,在盐、干旱胁迫下比较GUS酶活。 结果 鉴定出2段能够与ThbHLH1转录因子结合的DNA序列:分别为CCGAAA(LTRE1)和TGAC(WRKY710S)。 结论 在盐或干旱胁迫下,ThbHLH1通过与LTRE1或WRKY710S元件作用来激活基因表达。 Abstract:Objective To further study the mechanism of ThbHLH1 overexpression on improving the salt and drought tolerance of Tamarix hispida. Method The cis-acting elements recognized by ThbHLH1 were identified with the transcription factor-centered yeast one hybridization (TF-centered Y1H) method. To confirm the results of TF-centered Y1H, the effector construct 35S:ThbHLH1 was co-transformed with each reporter constructs pCAM-Cis into tobacco leaves with the particle bombardment method, and then the GUS activity was determined under salt or drought stress. Result ThbHLH1 could recognize LTRE1 (CCGAAA) and WRKY710S (TGAC) Cis-acting elements. Conclusion The results show that ThbHLH1 activates gene expression under salt or drought stress by interacting with LTRE1 or WRKY710S motifs. -
表 1 构建pGADT7-Rec2-ThbHLH1载体引物序列
Table 1. Primer sequences used in construction of the recombinant vector pGADT7-Rec2-ThbHLH1
引物名称
Primer name引物序列(5’-3’)
Sequence (5’-3’)Rec2-ThbHLH1 F AAGCAGTGGTATCAACGCAGAGTGGCCATTATGGCCCATGGCTAATAATCCAGGAG Rec2-ThbHLH1 R TCTAGAGGCCGAGGCGGCCGACATGTTATGAAAGGGGATTTGTC 注:_为pGADT7-Rec2载体的同源序列。 Note:_represents the homologous sequences of pGADT7-Rec2 vector. 表 2 构建随机DNA文库引物序列
Table 2. Primers used in constructing a random short DNA sequence insertion library
引物名称
Primer name引物序列(5’-3’)
Sequence (5’-3’)Y1 
Y2 
Y3 CGCGAACGCGTGAGCTC 注:_为Sma I线性化pHIS2载体两端的序列, 为随机DNA序列, N: A/G/C/T;Y: C/T Note:_represents the sequences on both ends of the Sma I linearized pHIS2 vector, represents the random DNA sequences, N: A/G/C/T;Y: C/T 表 3 构建酵母报告载体pHIS2-Cis引物序列
Table 3. Primer sequences used in constructing pHIS2-Cis yeast reporter vectors
引物名称
Primer name引物序列(5’-3’)
Sequence (5’-3’)pHIS2-LTRE1F AATTCCCGAAACCGAAACCGAAAGAGCT pHIS2-LTRE1R CTTTCGGTTTCGGTTTCGGG pHIS2-L1M1F AATTCAAGAAAAAGAAAAAGAAAGAGCT pHIS2-L1M1R CTTTCTTTTTCTTTTTCTTG pHIS2-L1M2F AATTCCCAAAACCAAAACCAAAAGAGCT pHIS2-L1M2R CTTTTGGTTTTGGTTTTGGG pHIS2-L1M3F AATTCCCGCCACCGCCACCGCCAGAGCT pHIS2-L1M3R CTGGCGGTGGCGGTGGCGGG pHIS2-L1M4F AATTCCCGAACCCGAACCCGAACGAGCT pHIS2-L1M4R CGTTCGGGTTCGGGTTCGGG pHIS2-L1M5F AATTCAAAAAAAAAAAAAAAAAAGAGCT pHIS2-L1M5R CTTTTTTTTTTTTTTTTTTG pHIS2-L1M6F AATTCCCGCCCCCGCCCCCGCCCGAGCT pHIS2-L1M6R CGGGCGGGGGCGGGGGCGGG pHIS2-L1M7F AATTCAAACCCAAACCCAAACCCGAGCT pHIS2-L1M7R CGGGTTTGGGTTTGGGTTTG pHIS2-WRKY710SF AATTCTGACTGACTGACGAGCT pHIS2-WRKY710SR CGTCAGTCAGTCAG pHIS2-WM1F AATTCCGACCGACCGACGAGCT pHIS2-WM1R CGTCGGTCGGTCGG pHIS2-WM2F AATTCTAACTAACTAACGAGCT pHIS2-WM2R CGTTAGTTAGTTAG pHIS2-WM3F AATTCTGCCTGCCTGCCGAGCT pHIS2-WM3R CGGCAGGCAGGCAG pHIS2-WM4F AATTCTGAATGAATGAAGAGCT pHIS2-WM4R CTTCATTCATTCAG pHIS2-WM5F AATTCCAACCAACCAACGAGCT pHIS2-WM5R CGTTGGTTGGTTGG pHIS2-WM6F AATTCTGCATGCATGCAGAGCT pHIS2-WM6R CTGCATGCATGCAG pHIS2-WM7F AATTCCACACACACACAGAGCT pHIS2-WM7R CTGTGTGTGTGTGG 注:_为粘性末端。Note: represents the sticky ends. 表 4 构建报告载体及效应载体引物序列
Table 4. Primer sequences used in constructing reporter and effector vectors
引物名称
Primer name引物序列(5’-3’)
Sequence (5’-3’)pCAM-LTRE1F AGCTTCCGAAACCGAAACCGAAAACCCTTCCTCTATATAAGGAAGTTCATTTCATTTGGAGAGAACACGGC pCAM-LTRE1R CATGGCCGTGTTCTCTCCAAATGAAATGAACTTCCTTATATAGAGGAAGGGTTTTCGGTTTCGGTTTCGGA pCAM-L1M7F AGCTTAAACCCAAACCCAAACCCACCCTTCCTCTATATAAGGAAGTTCATTTCATTTGGAGAGAACACGGC pCAM-L1M7R CATGGCCGTGTTCTCTCCAAATGAAATGAACTTCCTTATATAGAGGAAGGGTGGGTTTGGGTTTGGGTTTA pCAM-WRKY710SF AGCTTTGACTGACTGACACCCTTCCTCTATATAAGGAAGTTCATTTCATTTGGAGAGAACACGGC pCAM-WRKY710SR CATGGCCGTGTTCTCTCCAAATGAAATGAACTTCCTTATATAGAGGAAGGGTGTCAGTCAGTCAA pCAM-WM7F AGCTTCACACACACACAACCCTTCCTCTATATAAGGAAGTTCATTTCATTTGGAGAGAACACGGC pCAM-WM7R CATGGCCGTGTTCTCTCCAAATGAAATGAACTTCCTTATATAGAGGAAGGGTTGTGTGTGTGTGA pROKⅡ-ThbHLH1 F 
pROKⅡ-ThbHLH1 R 
注:_为粘性末端, 
Note:_ represents the sticky ends, 
表 5 ThbHLH1转录因子识别的插入序列分析
Table 5. Analysis of the insertion sequences bind to ThbHLH1
随机DNA插入序列
(5′-3′)Random DNA insertion
sequences (5′-3′)元件预测
Motif prediction参考文献
ReferencesCCCCCCGAAACGGG LTRE1: “CCGAAA” [22] CCCCGCTGACCGGG WRKY710S: “TGAC” [23] 注:_为文库的插入序列,左侧的CCC及右侧的GGG为载体侧翼序列,因为载体序列也可能和插入序列形成新的元件,所以要考虑左右两端各3个碱基的插入序列。 Note: _ represents the insertion sequences of the library, the left CCC and the right GGG are the vector flanking sequences, the insertion sequences of the three bases on the left and right sides should be considered, because the vector sequence may also form new motifs with the insertion sequences. -
[1] Peng H H, Shan W, Kuang J F, et al. Molecular characterization of cold-responsive basic helix-loop-helix transcription factors MabHLHs that interact with MaICE1 in banana fruit[J]. Planta, 2013, 238(5):937-953. doi: 10.1007/s00425-013-1944-7 [2] Liu W, Tai H, Li S, et al. bHLH122 is important for drought and osmotic stress resistance in Arabidopsis and in the repression of ABA catabolism[J]. New Phytologist, 2014, 201(4):1192-1204. doi: 10.1111/nph.12607 [3] Feller A, Machemer K, Braun E L, et al. Evolutionary and comparative analysis of MYB and bHLH plant transcription factor[J]. Plant Physiology, 2011, 66(1):94-116. [4] Robinson K A, Koepke J I, Kharodawala M, et al. A network of yeast basic helix-loop-helix interactions[J]. Nucleic Acids Research, 2000, 28(22):4460-4466. doi: 10.1093/nar/28.22.4460 [5] Toledo-Ortiz G, Huq E, Quail P H. The Arabidopsis basic/helix-loop-helix transcription factor family[J]. Plant Cell, 2003, 15(8):1749-1770. doi: 10.1105/tpc.013839 [6] Reguera M, Peleg Z, Blumwald E. Targeting metabolic pathways for genetic engineering abiotic stress-tolerance in crops[J]. Biochim Biophys Acta, 2012, 1819(2):186-194. doi: 10.1016/j.bbagrm.2011.08.005 [7] Heisler M G, Atkinson A, Bylstra Y H, et al. SPATULA, a gene that controls development of carpel margin tissues in Arabidopsis, encodes a bHLH protein[J]. Development, 2001, 128(7):1089-1098. [8] Ito S, Song Y H, Josephson-Day A R, et al. FLOWERING BHLH transcriptional activators control expression of the photoperiodic flowering regulator CONSTANS in Arabidopsis[J]. Proc Natl Acad Sci, USA, 2012, 109(9):3582-3587. doi: 10.1073/pnas.1118876109 [9] Zhao H, Wang X, Zhu D, et al. A single amino acid substitution in Ⅲf subfamily of basic helix-loop-helix transcription factor AtMYC1 leads to trichome and root hair patterning defects by abolishing its interaction with partner proteins in Arabidopsis[J]. Journal of Biological Chemistry, 2012, 287(17):14109-14121. doi: 10.1074/jbc.M111.280735 [10] Reymond M C, Brunoud G, Chauvet A, et al. A light-regulated genetic module was recruited to carpel development in Arabidopsis following a structural change to SPATULA[J]. Plant Cell, 2012, 24(7):2812-2825. doi: 10.1105/tpc.112.097915 [11] Leivar P, Tepperman J M, Cohn M M, et al. Dynamic antagonism between phytochromes and PIF family basic helixloop-helix factors induces selective reciprocal responses to light and shade in a rapidly responsive transcriptional network in Arabidopsis[J]. Plant Cell, 2012, 24(4):1398-1419. doi: 10.1105/tpc.112.095711 [12] Zhang L Y, Bai M Y, Wu J, et al. Antagonistic HLH/bHLH transcription factors mediate brassinosteroid regulation of cell elongation and plant development in rice and Arabidopsis[J]. Plant Cell, 2009, 21(12):3767-3780. doi: 10.1105/tpc.109.070441 [13] Jiang Y Q, Yang B, Deyholos M K. Functional characterization of the Arabidopsis bHLH92 transcription factor in abiotic stress[J]. Molecular Genetics and Genomics, 2009, 282(5):503-516. doi: 10.1007/s00438-009-0481-3 [14] Chinnusamy V, Ohta M, Kanrar S, et al. ICE1:a regulator of cold-induced transcriptome and freezing tolerance in Arabidopsis[J]. Genes & Development, 2003, 17(8):1043-1054. [15] Nakata M, Mitsuda N, Herde M, et al. A bHLH-type transcription factor, ABA-INDUCIBLE BHLH-TYPE TRANSCRIPTION FACTOR/JA-ASSOCIATED MYC2-LIKE1, acts as a repressor to negatively regulate jasmonate signaling in Arabidopsis[J]. Plant Cell, 2013, 25(5):1641-1656. doi: 10.1105/tpc.113.111112 [16] Seo J S, Joo J, Kim M J, et al. OsbHLH148, a basic helix-loop-helix protein, interacts with OsJAZ proteins in a jasmonate signaling pathway leading to drought tolerance in rice[J]. Plant Journal, 2011, 65(6):907-921. doi: 10.1111/tpj.2011.65.issue-6 [17] Zhou J, Li L, Wang J L, et al. Basic helix-loop-helix transcription factor from wild rice (OrbHLH2) improves tolerance to salt-and osmotic stress in Arabidopsis[J]. Journal of Plant Physiology, 2009, 166(12):1296-1306. doi: 10.1016/j.jplph.2009.02.007 [18] Yang T, Yao S, Hao L, et al. Wheat bHLH-type transcription factor gene TabHLH1 is crucial in mediating osmotic stresses tolerance through modulating largely the ABA-associated pathway[J]. Plant Cell Rep, 2016, 35(11):2309-2323. doi: 10.1007/s00299-016-2036-5 [19] Ji X Y, Nie X G, Liu Y J, et al. A bHLH gene from Tamarix hispida improves abiotic stress tolerance by enhancing osmotic potential and decreasing reactive oxygen species accumulation[J]. Tree Physiology, 2016, 36(2):193-207. [20] Ji X Y, Wang L Q, Nie X G, et al. A novel method to identify the DNA motifs recognized by a defined transcription factor[J]. Plant Molecular Biology, 2014, 86(4-5):367-380. doi: 10.1007/s11103-014-0234-5 [21] Jefferson R A, Kavanagh T A, Bevan M W. GUS fusions:betaglucuronidase as a sensitive and versatile gene fusion marker in higher plants[J]. EMBO Journal, 1987, 6(13):3901-3907. doi: 10.1002/embj.1987.6.issue-13 [22] Dunn M A, White A J, Vural S, et al. Identification of promoter elements in a low-temperature-responsive gene (blt4.9) from barley (Hordeum vulgare L.)[J]. Plant Molecular Biology, 1998, 38(4):551-564. doi: 10.1023/A:1006098132352 [23] Matton D P, Prescott G, Bertrand C, et al. Identification of cis-acting elements involved in the regulation of the pathogenesis-related gene STH-2 in potato[J]. Plant Molecular Biology, 1993, 22(2):279-291. doi: 10.1007/BF00014935 [24] Chen Y Y, Li M Y, Wu X J, et al. Genome-wide analysis of basic helix-loop-helix family transcription factors and their role in responses to abiotic stress in carrot[J]. Molecular Breeding, 2015, 35: 125. [25] 杨鹏程, 周波, 李玉花.植物花青素合成相关的bHLH转录因子[J].植物生理学报, 2012, 48(8):747-758. [26] 光杨其, 宋桂成, 张金凤, 等. 1个新棉花bHLH类基因GhbHLH130的克隆及表达分析[J].棉花学报, 2014, 26(4):363-370. doi: 10.3969/j.issn.1002-7807.2014.04.012 [27] Xue G P. An AP2 domain transcription factor HvCBF1 activates expression of cold-responsive genes in barley through interaction with a (G/a)(C/t)CGAC motif[J]. Biochim Biophys Acta, 2002, 1577:63-72. doi: 10.1016/S0167-4781(02)00410-4 [28] Feng X M, Zhao Q, Zhao L L, et al. The cold-induced basic helix-loop-helix transcription factor gene MdCIbHLH1 encodes an ICE-like protein in apple[J]. BMC Plant Biol, 2012, 12: 22. [29] Huang X S, Wang W, Zhang Q, et al. A basic helix-loop-helix transcription factor, PtrbHLH, of Poncirus trifoliata confers cold tolerance and modulates peroxidase-mediated scavenging of hydrogen peroxide[J]. Plant Physiology, 2013, 162(2):1178-1194. doi: 10.1104/pp.112.210740 -




 下载:
下载: