-
油菜素甾醇类物质(Brassinosteroids, BRs)是植物中一类多羟基甾醇类化合物,油菜素内酯(Brassinolide, BL)是植物中最早被鉴定出的甾醇类激素,在植物整个生长发育和响应胁迫条件中都发挥了重要作用[1]。BL的生物合成受到多种酶的调控,其中,类固醇22-α羟化酶(DWF4)是最重要限速酶之一,DWF4的含量直接影响到高等植物体内油菜素内酯的合成量[2-3]。DWF4最先在拟南芥(Arabidopsis thaliana L.)的dwarf4突变体中被鉴定出来[4],该基因编码的蛋白酶属于细胞色素P450单加氧酶(CYP90B1)。目前,BL已在拟南芥和水稻等模式植物中广泛研究,尤其在其合成调控机制和适应胁迫调节方面较为深入。过量表达AtDWF4基因可以显著提高拟南芥分蘖数量、果荚个数及种子的产量[5],而且过表达AtDWF4可以打破由脱落酸引起的抑制种子萌发的现象,从而提高植株抵抗低温胁迫能力[6]。过量表达ZmDWF4可以使玉米叶片变得狭长,叶柄和花茎节间变长、株高增加、果实产量提高、花期提前、抗盐和水分胁迫能力增强[7]。
竹子是禾本科竹亚科一类植物,因其具有生长迅速,且融经济、生态价值为一体等诸多特征而倍受关注。关于竹子的生长发育及其适应性已有大量的研究报道,涉及栽培生理、植物化学、形态解剖、分子生物学等多个方面,尤其是毛竹基因组草图的公布[8]和基因组数据库[9]的建立与共享,极大促进了竹子基础研究的发展。虽然关于竹子内源激素(吲哚乙酸、玉米素、赤霉素、油菜素内酯和脱落酸)在笋期的含量[10-14]、在笋芽中的比例[15]已有相关研究报道,然而关于竹子BL生物合成的相关研究尚未有报道。本研究以重要经济竹种毛竹(Phyllostachys edulis Carr.)为实验材料,通过分离毛竹中DWF4同源基因,在对其基因结构特征分析的基础上,对其蛋白保守结构域,进化关系,组织表达模式以及在高盐、干旱、低温和强光胁迫条件下的表达量变化规律进行了分析,以期为研究该基因在响应不同胁迫的调节机制奠定基础,同时也为深入了解竹子中BL的生物合成及其调控作用提供参考。
-
取购买自广西桂林的毛竹种子于实验室条件下培养,培养基质为泥炭:蛭石=7:3,温度28℃,相对湿度50%,光照150 μmol·m-2·s-1,光周期为16 h光照/8 h黑暗,加施B5营养液。取半年生实生苗用于实验处理。
选取长势一致的盆栽毛竹实生苗,从培养基质中整株取出,清水冲洗干净。盐处理:毛竹实生苗根浸泡在400 mmol·L-1的NaCl溶液中,分别取处理1 h(叶片开始出现卷曲)、2、4和6 h(完全卷曲)的叶片;干旱处理:直接将裸根竹苗置于实验室条件下处理,取处理1、2 h(叶片开始出现卷曲)、4和8 h(完全卷曲)的叶片样本;低温处理:将盆栽毛竹实生苗于4℃下进行处理,分别取处理1、2、4、8 h的叶片样本;强光处理:将盆栽毛竹实生苗于1 200 μmol·m-2·s-1强光下培养,分别取1、2、4和8 h叶片样本。各处理均以正常生长的毛竹叶片为对照。另外,取正常生长的盆栽毛竹的根、茎、完全展开叶片、未完全展开叶片、叶鞘和笋样品。所有样品于液氮中速冻,并存放于-80℃冰箱中用于RNA提取。
-
利用改良TRIzol法[16]提取毛竹各样本的总RNA,使用DNase I(Promega, 美国)去除残余DNA,并使用反转录试剂盒(Promega, 美国)合成cDNA第一条链。
-
以拟南芥中DWF4基因(AT3G50660)为目的序列,在毛竹数据库BambooGDB (http://www.bamboogdb.org/)中Blast得到DWF4基因同源序列。使用Primer Premier 5.0软件设计开放阅读框引物(PeDWF4-F和PeDWF4-R),并由上海生工技术有限公司合成,引物基本信息见表 1。
表 1 PCR扩增所用引物序列
Table 1. Primer sequences used for PCR
引物名称
Primer name序列(5′→3′)
Sequence (5′→3′)用途
ApplicationPeDWF4-F ATGGC CTCCATAACCAGCGAG 开放阅读框扩增 PeDWF4-R TTACTCTTCCTCCTGTGCAATTCTATG Open reading frame amplification PeDWF4-F1 GGCTAGGCAACGTGGTCAGG 半定量引物 PeDWF4-R1 TTGTTCTTC CATCTCCAAGGGTT Primers for semi-quantitative analysis PeActin-F GATCTTGCTGGGCGTGACCTC 半定量内参引物 PeActin-R CCATCGGGCATCTCGTAGC Internal reference primers for semi-quantitative analysis PeDWF4-qF GGCTAGGCAACGTGGTCAGG 定量引物 PeDWF4-qR TTGTTCTTCCATCTCCAAGGGTT Primers for quantitative analysis PeNTB-F TCTTGTTTGACACCGAAGAGGAG 定量内参引物 PeNTB-R AATAGCTGTCCCTGGAGGAGTTT Internal reference primers for quantitative analysis 毛竹分别以叶片cDNA和基因组DNA为模板,使用Prime STAR聚合酶扩增目的基因的编码区序列及其基因组序列。20 μL PCR反应体系:10 × Prime STAR Buffer 2 μL,Prime STAR酶(2.5 U·μL-1)0.2 μL,cDNA(40 ng·mL-1)1 μL,dNTP(dATP、dTTP、dCTP和dGTP各2.5 mmol·L-1)1.6 μL,PeDWF4-F和PeDWF4-R(10 μmol·L-1)各1 μL,ddH2O 3.2 μL。反应条件为:98℃,4 min;98℃ 30 s,64℃ 30 s,72℃ 90 s,34个循环;72℃ 10 min。回收PCR产物,并由上海生工技术有限公司测序。
使用GSDS(http://gsds.cbi.pku.edu.cn/)分析基因结构,用ProtParam (http://web.expasy.org/ protparam/)在线软件分析蛋白的基本理化性质,用ClustalW2[17]比对同源序列并分析保守结构域[18],使用N-J法构建基于DWF4蛋白序列的系统发育进化树。
-
从NCBI的数据库(Short Read Archive)中下载毛竹7个不同组织的转录组数据[19],利用PeDWF4基因的表达值绘制基因表达热图(http://www.chibi.ubc.ca/matrix2png/)。
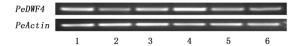
使用引物PeDWF4-F1和PeDWF4-R1对毛竹根、茎、完全展叶片、未完全展开叶片、叶鞘和笋6个组织中表达量进行分析,同时以毛竹PeActin基因为内参进行表达量分析[20]。
-
根据PeDWF4基因的特异序列设计定量引物(PeDWF4-qF和PeDWF4-qR),利用qTOWER2.2(Analytik Jena, 德国)PCR仪进行定量分析PCR,每个反应4次重复。反应体系为:LightCycler® 480 SYBR Green I Master Mix(Roche, 美国)5.0 μL,正、反向引物各0.2 μL,cDNA 0.8 μL,ddH2O 3.8 μL。反应程序:95℃ 6 min;95℃ 10 s,63℃ 12 s,共45个循环。同时以毛竹PeNTB作为内参基因[21],引物为PeNTB-F和PeNTB-R(表 1)。3次生物学实验后,利用2-ΔΔCT法[22]分析基因的表达规律。
-
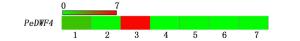
通过在BambooGDB比对得到毛竹中DWF4同源序列PH01001012G0570。以毛竹cDNA为模板,用PeDWF4-F和PeDWF4-R扩增获得的基因编码区序列与PH01001012G0570序列完全一致,该基因全长为1 503 bp,编码的蛋白长度为500 aa,分子量为57.36 kDa,理论等电点为9.14呈碱性,不稳定系数为57.78,平均亲水系数(GRAVY)为-0.217,脂肪系数为89.56,属于细胞色素P450家族的一种单加氧酶,命名为PeDWF4;同时,PeDWF4编码区对应的基因组序列为6 149 bp,由7个内含子和8个外显子组成(图 1),其转录过程中内含子剪接属于GT-AG类型[23]。
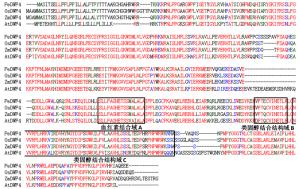
利用ClustalW2对PeDWF4编码蛋白和水稻、玉米、拟南芥等模式植物氨基酸同源序列进行比对分析,结果表明:它们都含有细胞色素P450单加氧酶特有的血红素结合域A、类固醇结合结构域B和C等(图 2),进而推测PeDWF4编码的蛋白质在生长发育过程中可能发挥着与其他物种中DWF4相似的功能,同时也验证了PeDWF4是细胞色素P450家族成员。
构建基于毛竹、二穗短柄草(Brachypodium distachyon L.)、节节麦(Aegilops tauschii Coss.)、大麦(Hordeum vulgare L.)、小麦(Triticum aestivum L.)、水稻(Oryza sativa L.)、玉米(Zea mays L.)、粟(Setaria italica L.)、高粱(Sorghum bicolor L.)、葡萄(Vitis vinifera L.)、大豆(Glycine max L.)、拟南芥的DWF4同源基因编码氨基酸序列的进化树(图 3)。
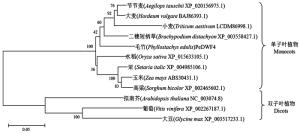
图 3 基于DWF4氨基酸序列的系统进化树
Figure 3. Phylogenetic tree constructed on the basis of DWF4 amino acid sequences
结果显示:DWF4基因编码蛋白的进化在高等植物中分支成单子叶植物和双子叶植物2个类群,而作为禾本科竹亚科的毛竹与水稻、二穗短柄草、玉米等禾本科植物的DWF4蛋白同属于一个系统发育分支,亲缘关系较近,而与拟南芥、葡萄和大豆等双子叶植物的亲缘关系较远,这与形态学分类相一致。
-
利用毛竹RNA-Seq表达谱数据,分析PeDWF4基因的组织表达模式。结果(图 4)显示:PeDWF4基因呈现组成型表达,但差异明显,其中,叶片中表达丰度最高,根中表达次之,而在地下茎、盛花期花序、50 cm笋、早花期花序和20 cm笋中表达量均较低。
RT-PCR结果显示:PeDWF4在所检测的毛竹6个组织中均有表达,且差异明显(图 5),其中,叶片中的表达量最高,且未完全展开叶高于完全展开叶,这可能与叶片区域成熟有关,根中表达量次之。该结果与RNA-Seq表达谱相符合,均证明PeDWF4在叶片中表达量最高。因此,选取新鲜未完全展开的毛竹叶片作为实验材料进行处理并分析其表达模式。
-
DWF4基因在提高植物抗逆性,响应高盐、干旱、低温和强光胁迫等环境胁迫方面发挥着重要作用[7]。在高盐、干旱、低温和强光等胁迫条件下,毛竹叶片中PeDWF4表达模式因处理条件而呈现不同的变化,但整体趋势一致,相对表达量均呈先上升后下降(图 6)。
NaCl和干旱胁迫条件下,叶片中PeDWF4的表达量随处理时间均为先诱导后抑制(图 6A、B)。PeDWF4表达量在NaCl处理2 h达到最高,约为对照的3.5倍,随后表达量随处理时间推移迅速下降,在处理6 h时达到最低,此时叶片已完全卷曲,PeDWF4的表达量仅为对照的20%(图 6A)。在干旱胁迫条件下,处理1 h后PeDWF4的表达量达到最高,约为对照的2倍,随后表达量随处理时间持续下降,8 h后达到最低,约为对照的60%(图 6B)。
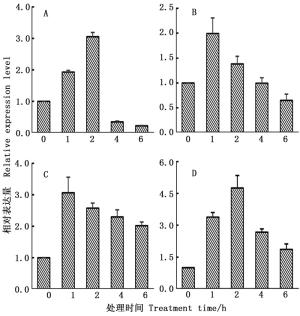
图 6 不同胁迫条件下PeDWF4基因在毛竹叶片中的表达量
Figure 6. Expression of PeDWF4 in leaves under different stresses of Ph. edulis
低温和强光胁迫条件下,PeDWF4在叶片中均表现为诱导表达,表达量随时间推移表现为先上升后下降(图 6C、D)。强光处理下,PeDWF4表达量在处理2 h后达到最高,约为对照的4.5倍,之后随处理时间延长表达量呈下降趋势,至8 h时基因表达量仍为对照的2倍(图 6D)。低温条件下,与强光胁迫处理结果表达量变化趋势基本一致,PeDWF4表达量表现为先上升后下降的变化趋势,处理1 h后达到最高,约为对照的3倍,之后随时间推移持续下降,至8 h时基因表达量最低,约为对照的2倍(图 6C)。
-
BL作为一种微量高效的植物激素,具有参与调节细胞的伸长和分裂[24]、促进根的伸长和种子的萌发[25]、加速植物叶片细胞的衰老[26]、影响高等植物的开花周期[27]、调节植物的光形态建成[28]、増强光合作用强度[29]、维管束分化[30]、植物的育性、内源激素平衡[29]等作用。DWF4是BL合成通路中的关键酶基因,其表达量与BL合成量存在正相关关系[31]。因此,通过对毛竹PeDWF4基因进行研究,以期为深入了解PeDWF4在竹子BL生物合成中的作用提供参考。序列比对结果显示,PeDWF4编码的蛋白包含细胞色素P450单加氧酶特有保守结构域[18],且与水稻中的OsDWF4相似性最高,表明PeDWF4编码的蛋白在进化上的相对保守,意味着其可能具有类固醇22-α羟化酶的功能。基因表达组织模式分析表明,PeDWF4为组成型表达,其中, 表达量最高出现在叶片中,因此, 推测其可能在叶片中发挥主要功能。未完全展开叶片中PeDWF4的表达量高于完全展开叶片,这可能是由于随着叶片的成熟,形态建成完成,促进生长的关键酶基因表达量降低引起的[28, 30]。
高盐、干旱、低温和强光是植物最常遇到的非生物胁迫,已有研究表明,过量表达胡杨DWF4基因的烟草能够提高其对盐和干旱的抵抗能力[6, 32], 过表达AtDWF4能够提高转基因荠菜(Brassica juncea)植株的抗寒能力,有助于低温(4℃)处理后的恢复生长[33]。PeDWF4和AtDWF4编码的蛋白具有较高的一致性(91%),意味着它们可能具有相似功能。NaCl和干旱胁迫条件下,PeDWF4基因在叶中表达量随处理时间先上升后下降,这可能是由于PeDWF4通过提高自身表达量促进BL的合成,进而提高植物抗盐胁迫的能力[7],但随着处理时间延长叶片逐步受到伤害,其生理活性降低,导致该基因的表达量降低。在低温和强光胁迫条件下,叶片中PeDWF4表达量均表现为诱导模式,变化趋势为先上升后下降,在处理初期通过提高表达量来响应低温和强光胁迫,但低温和强光对毛竹的胁迫程度有限,随着时间推移植物通过调节自身代谢逐渐适应外界变化环境,基因表达量故逐步下降接近平稳水平。因此,推测PeDWF4在毛竹响应胁迫过程中发挥了一定的调节作用,可能是通过上调PeDWF4基因的表达量,增加DWF4酶含量促进C-22羟基化催化反应发生,使油菜素内酯生物合成量增加,进一步通过油菜素内酯调控胁迫相关基因和与其他植物激素的相互作用,来提高对环境胁迫的抵御能力。
-
通过同源基因克隆的方法获得了毛竹PeDWF4基因,该基因编码的碱性蛋白长度为500 aa,具有细胞色素P450家族的血红素结合域A、类固醇结合结构域B和C等3个特定结构域,属于单加氧酶。毛竹中PeDWF4呈现组成型表达,在叶片中表达量最高。在高盐、干旱、寒冷和强光胁迫条件下,毛竹叶片中PeDWF4的表达均呈先上升后下降的应答趋势。本研究为解释PeDWF4基因功能和其响应胁迫机理提供参考依据。
毛竹PeDWF4基因克隆及表达模式分析
Cloning and Expression Pattern Analysis of PeDWF4 Gene in Moso Bamboo (Phyllostachys edulis)
-
摘要:
目的 通过对毛竹PeDWF4基因结构特点和表达特征的研究,揭示其在响应逆境胁迫过程中的作用。 方法 采用同源序列比对的方法,从毛竹基因组数据库中获得DWF4同源基因信息并克隆,通过生物信息学方法分析该基因的结构、理化特征,以及基因编码蛋白的保守结构域、进化关系等,应用RT-PCR技术分析基因在毛竹不同组织中的表达情况,使用实时荧光定量PCR技术分别分析高盐、干旱、低温和强光等胁迫条件下该基因在叶片中的表达模式。 结果 从毛竹中克隆获得1个DWF4同源基因PeDWF4,编码区长度为1 503 bp,对应的基因组序列为6 149 bp,包含8个外显子和7个内含子,内含子完全符合GT-AG剪接原则。PeDWF4编码1个500 aa的碱性蛋白,属于细胞色素P450家族的单加氧酶。组织特异性表达分析表明,PeDWF4在毛竹不同组织中均检测到表达,其中,叶片中表达丰度最高,其次是根,茎、叶鞘和笋中的表达量较低。在NaCl(400 mmol·L-1)和干旱胁迫条件下,叶片中PeDWF4的表达均先受到诱导,后受到抑制,其中,NaCl处理下,表达量在2 h时达到最高(为对照的3.5倍),6 h时最低(为对照的20%);干旱处理下,表达量在1 h时达到最高(为对照的2倍),8 h时最低(为对照的60%)。强光(1 200 μmol·m-2·s-1)和低温(4℃)胁迫均诱导PeDWF4的表达,其中,强光处理2 h时表达量达到最高(为对照的4.5倍),随后降低,8 h时仍为对照的2倍;低温处理下,叶片中PeDWF4表达量在1 h时达到最高(约为对照的3倍),随后持续下降,8 h时仍为对照的2倍。 结论 从毛竹中克隆了BL生物合成关键限速酶基因PeDWF4,该基因在毛竹中呈现组成型表达,在叶片中的表达受到NaCl、干旱、低温和强光等非生物胁迫的影响,基因表达的变化表明PeDWF4可能有助于毛竹适应逆境胁迫。 Abstract:Objective This study aims to provide reference for revealing the role of steroid 22-alpha hydroxylase in moso bamboo (PeDWF4) in the response to abiotic stresses, based on the analysis of its gene structural characteristics and expression patterns. Method The method of homologous sequence comparison was used to isolate the homologous gene of DWF4 in moso bamboo with the information in BambooGDB. Bioinformatic method was used to analyze the gene structure, the basic physical and chemical characteristics, the conservative domains in the protein encoded by the gene, and the evolutionary relationships, etc. Besides, RT-PCR was applied for the gene expression analysis in different bamboo tissues and real-time fluorescent quantitative PCR (qRT-PCR) was conducted to find the expression patterns of the gene in leaf blades under the stresses of high salt, drought, low temperature and high light, respectively. Result PeDWF4, a homologous gene of DWF4, was isolated from moso bamboo, whose open reading frame is 1 503 bp, and the corresponding genome sequence is 6 149 bp containing 8 exons and 7 introns. The introns completely conformed to the principle of GT-AG splicing. PeDWF4 encoded an alkaline protein with 500 aa, belonging to the single oxygenases of cytochrome P450 family. Tissue specific expression analysis showed that PeDWF4 was detected in all tissues of bamboo roots, stems, fully expanded leaf blades, not fully expanded leaf blades, leaf sheaths and shoots with different levels, among which the highest one was in leaf blades, followed by roots, while those in stems, sheaths and shoots were relatively lower. In accordance with the transcriptome data, PeDWF4 had the highest expression level in leaf blades. Both under NaCl (400 mmol·L-1) and drought stresses, the expression of PeDWF4 in leaf blade showed a trend of induction and then suppression. Under NaCl stress, it was upregulated to 3.5 times of the control after 2 hours, then decreased gradually with the prolonged processing and reached to nearly only 20% of the control after 6 hours. That under drought stress was upregulated to 2 times of the control after 1 hour, and decreased to 60% of the control after 8 hours. The expression of PeDWF4 was induced by both high light (1 200 μmol·m-2·s-1) and low temperature (4℃) stresses. Under high light stress, it was upregulated to 4.5 times of the control after 2 hours, then decreased gradually with the prolonged processing and reached to 2 times of the control after 8 hours. That under low temperature stress was upregulated to 3 times of the control after 1 hour, and then decreased gradually and reached to 2 times of the control after 8 hours. Conclusion PeDWF4, a DWF4 homologous gene is isolated from moso bamboo, which belongs to the single oxygenase gene of cytochrome P450 family. PeDWF4 is constitutively expressed in moso bamboo. The expression changes indicate that PeDWF4 is involved in response to NaCl, drought, low temperature and high light stresses, which might be helpful to improve the ability of moso bamboo to withstand abiotic stresses. -
Key words:
- Phyllostachys edulis
- / steroid 22-alpha hydroxylase
- / DWF4
- / abiotic stresses
- / gene expression
-
表 1 PCR扩增所用引物序列
Table 1. Primer sequences used for PCR
引物名称
Primer name序列(5′→3′)
Sequence (5′→3′)用途
ApplicationPeDWF4-F ATGGC CTCCATAACCAGCGAG 开放阅读框扩增 PeDWF4-R TTACTCTTCCTCCTGTGCAATTCTATG Open reading frame amplification PeDWF4-F1 GGCTAGGCAACGTGGTCAGG 半定量引物 PeDWF4-R1 TTGTTCTTC CATCTCCAAGGGTT Primers for semi-quantitative analysis PeActin-F GATCTTGCTGGGCGTGACCTC 半定量内参引物 PeActin-R CCATCGGGCATCTCGTAGC Internal reference primers for semi-quantitative analysis PeDWF4-qF GGCTAGGCAACGTGGTCAGG 定量引物 PeDWF4-qR TTGTTCTTCCATCTCCAAGGGTT Primers for quantitative analysis PeNTB-F TCTTGTTTGACACCGAAGAGGAG 定量内参引物 PeNTB-R AATAGCTGTCCCTGGAGGAGTTT Internal reference primers for quantitative analysis -
[1] Grove M D, Spencer G F, Rohwedder W K, et al. Brassinolide a plant growth-promoting steroid isolated from Brassica napus pollen[J]. Nature, 1979, 281(5728):216-217. doi: 10.1038/281216a0 [2] Palatnik J F, Allen E, Wu X, et al. Control of leaf morphogenesis by microRNAs[J]. Nature, 2003, 425(6955):257-263. doi: 10.1038/nature01958 [3] Koyama T, Furutani M, Tasaka M, et al. TCP transcription factors control the morphology of shoot lateral organs via negative regulation of the expression of boundary-specific genes in Arabidopsis[J]. The Plant Cell Online, 2007, 19(2):473-484. doi: 10.1105/tpc.106.044792 [4] Azpiroz R, Wu Y, Lo Cascio J C, et al. An Arabidopsis brassinosteroid-dependent mutant is blocked in cell elongation[J]. The Plant Cell, 1998, 10(2):219-230. doi: 10.1105/tpc.10.2.219 [5] Choe S, Fujioka S, Noguchi T, et al. Overexpression of DWARF4 in the brassinosteroid biosynthetic pathway results in increased vegetative growth and seed yield in Arabidopsis[J]. Plant Journal, 2001, 26(6):573-582. doi: 10.1046/j.1365-313x.2001.01055.x [6] Divi U K, Rahman T, Krishna P. Brassinosteroid-mediated stress tolerance in Arabidopsis shows interactions with abscisic acid, ethylene and salicylic acid pathways[J]. BMC Plant Biology, 2010, 10(1):151. doi: 10.1186/1471-2229-10-151 [7] Liu T, Zhang J, Wang M, et al. Expression and functional analysis of ZmDWF4, an ortholog of Arabidopsis DWF4 from maize (Zea mays L.)[J]. Plant Cell Report, 2007, 26(12):2091-2099. doi: 10.1007/s00299-007-0418-4 [8] Peng Z H, Lu Y, Li L B, et al. The draft genome of the fast-growing non-timber forest species moso bamboo (Phyllostachys heterocycla)[J]. Nature Genetic, 2013, 45(4):456-461. doi: 10.1038/ng.2569 [9] Zhao H., Peng Z, Fei B, et al. BambooGDB:a bamboo genome database with functional annotation and an analysis platform[J]. Database, 2014, 2014(10):bau006. [10] 张春玲.毛竹笋-竹生长发育过程系统分析与生长素相关基因研究[D].北京: 中国林业科学研究院, 2014: 5-6. [11] Wang H Y, Cui K, He C Y, et al. Endogenous hormonal equilibrium linked to bamboo culm development[J]. Genetics and Molecular Research, 2015, 14(3):11312-11323. doi: 10.4238/2015.September.22.25 [12] 方楷, 杨光耀, 杨清培, 等.毛竹成竹过程中内源激素动态变化[J].江西农业大学学报, 2011, 33(6):1107-1111. doi: 10.3969/j.issn.1000-2286.2011.06.014 [13] 丁兴萃.毛竹笋体生长发育过程中内源激素的动态分析[J].竹子研究汇刊, 1997, 16(2):53-62. [14] 崔凯.毛竹茎秆快速生长的机理研究[D].北京: 中国林业科学研究院, 2011: 5-6. [15] 黄坚钦, 刘力, 章滨森, 等.雷竹地下鞭侧芽内源激素的动态变化研究[J].林业科学, 2002, 38(3):38-41. doi: 10.3321/j.issn:1001-7488.2002.03.007 [16] 高志民, 范少辉, 高健, 等.基于CTAB法提取毛竹基因组DNA的探讨[J].林业科学研究, 2006, 19(6):725-728. doi: 10.3321/j.issn:1001-1498.2006.06.009 [17] Thompson J D, Higgins D G, Gibson T J. CLUSTAL W:improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice[J]. Nucleic Acids Research, 1994, 22(22):4673-4680. doi: 10.1093/nar/22.22.4673 [18] Nebert D W, Gonzalez F J. P450 genes:structure, evolution, and regulation[J]. Annual Review of Biochemistry, 1987, 56(1):945-993. doi: 10.1146/annurev.bi.56.070187.004501 [19] Zhao H S, Dong L L, Sun H Y, et al. Comprehensive analysis of multitissue transcriptome data and the genome-wide investigation of GRAS family in Phyllostachys edulis[J]. Scientific Reports, 2016, 6:27640. doi: 10.1038/srep27640 [20] Sun H, Li L, Lou Y, et al. Cloning and preliminary functional analysis of PeUGE gene from moso bamboo (Phyllostachys edulis)[J]. DNA and Cell Biology, 2016, 35(11):706-714. doi: 10.1089/dna.2016.3389 [21] Fan C, Ma J, Guo Q, et al. Selection of reference genes for quantitative real-time PCR in bamboo (Phyllostachys edulis)[J]. PLoS One, 2013, 8(2):e56573. doi: 10.1371/journal.pone.0056573 [22] Livak K J, Schmittgen T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCt method[J]. Methods, 2001, 25(4):402-408. doi: 10.1006/meth.2001.1262 [23] Moore M J, Query C C, Sharp P A. Splicing of precursors to mRNA by the spliceosome[M]//Gesteland R F, Atkins J F. The RNA world Cold Spring Harbor:Cold Spring Harbor laboratory Press, 1993:303-357. [24] Clouse S D, Zurek D. Molecular analysis of brassinolide action in plant growth and development[M]//Cutler H G, Yokota T, Adam G. Brassinosteroids:Chemistry, Bioactivity & Applications, American Chemical Society, Washington DC, 1991:122-140. [25] Domagalskam M A, Schomburg F M, Amasino R M, et al. Attenuation of brassinosteroid signaling enhances FLC expression and delays flowering[J]. Development, 2007, 134(15):2841-2850. doi: 10.1242/dev.02866 [26] Choe S, Chung Y. The regulation of brassinosteroid biosynthesis in Arabidopsis[J]. Critical Reviews in Plant Sciences, 2013, 32(6):396-410. doi: 10.1080/07352689.2013.797856 [27] Hou X, Hu W W, Shen L, et al. Global identification of DELLA target genes during Arabidopsis flower development[J]. Plant Physiology, 2008, 147(3):1126-1142. doi: 10.1104/pp.108.121301 [28] Nakamoto D, Ikeura A, Asami T, et al. Inhibition of brassinosteroid biosynthesis by either a dwarf4 mutation or a brassinosteroid biosynthesis inhibitor rescues defects in tropic responses of hypocotyls in the Arabidopsis mutant nonphototropic hypocotyl4[J]. Plant Physiology, 2006, 141(2):456-464. doi: 10.1104/pp.105.076273 [29] Sasse J M. Physiological actions of brassinosteroids:An update[J]. Journal of Plant Growth Regulation, 2003, 22(4), 276-288. doi: 10.1007/s00344-003-0062-3 [30] Nomura T, Sato T, Bishop J, et al. Accumulation of 6-deoxocathasterone and 6-deoxocastasterone in Arabidopsis, pea and tomato is suggestive of common rate-limiting steps in bassinosteroid biosynthesis[J]. Phytochemistry, 2001, 57(2):171-178. doi: 10.1016/S0031-9422(00)00440-4 [31] Asami T, Mizutani M, Shimada Y, et al. Triadimefon, a fungicidal triazole-type P450 inhibitor, induces brassinosteroid deficiency-like phenotypes in plants and binds to DWF4 protein in the brassinosteroid biosynthesis pathway[J]. Biochemical Journal, 2003, 369(1):71-76. doi: 10.1042/bj20020835 [32] 王阳, 陈永富, 高永峰, 等.异源过表达胡杨PeDWF4基因提高烟草对非生物胁迫的耐性[J].基因组学与应用生物学, 2017(10):4242-4249. [33] 兰彩耘.超量表达AtDWF4基因对芥菜生长发育及抗寒性的影响[D].昆明: 西南大学, 2016. -




 下载:
下载: